Abstract
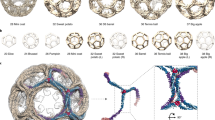
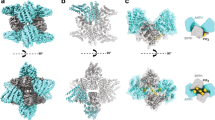
Clathrin is a triskelion-shaped cytoplasmic protein that polymerizes into a polyhedral lattice on intracellular membranes to form protein-coated membrane vesicles. Lattice formation induces the sorting of membrane proteins during endocytosis and organelle biogenesis by interacting with membrane-associated adaptor molecules1. The clathrin triskelion is a trimer of heavy-chain subunits (1,675 residues), each binding a single light-chain subunit, in the hub domain (residues 1,074–1,675). Light chains negatively modulate polymerization so that intracellular clathrin assembly is adaptor-dependent2. Here we report the atomic structure, to 2.6 Å resolution, of hub residues 1,210–1,516 involved in mediating spontaneous clathrin heavy-chain polymerization and light-chain association3,4. The hub fragment folds into an elongated coil of α-helices, and alignment analyses reveal a 145-residue motif that is repeated seven times along the filamentous leg and appears in other proteins involved in vacuolar protein sorting. The resulting model provides a three-dimensional framework for understanding clathrin heavy-chain self-assembly, light-chain binding and trimerization.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schmid, S. L. Clathrin-coated vesicle formation and protein sorting: an integrated process. Annu. Rev. Biochem. 66, 511–548 (1997).
Ybe, J. A.et al. Clathrin self-assembly is regulated by three light chain residues controlling the formation of critical salt bridges. EMBO J. 17, 1297–1303 (1998).
Näthke, I. S.et al. Folding and trimerization of clathrin subunits at the triskelion hub. Cell 68, 899–910 (1992).
Liu, S. H., Wong, M. L., Craik, C. S. & Brodsky, F. M. Regulation of clathrin assembly and trimerization defined using recombinant triskelion hubs. Cell 83, 257–267 (1995).
Smith, C. J., Grigorieff, N. & Pearse, B. M. F. Clathrin coats at 21 Å resolution: a cellular assembly designed to recycle multiple membrane receptors. EMBO J. 17, 4943–4953 (1998).
Chothia, C., Levitte, M. & Richardson, D. Helix to helix packing in proteins. J. Mol. Biol. 145, 215–250 (1981).
Raag, R., Appelt, K., Xuong, N. H. & Banaszak, L. Structure of the lamprey yolk lipid–protein complex lipovitellin–phosvitin at 2.8 Å resolution. J. Mol. Biol. 200, 553–569 (1988).
Thunnissen, A.-M.et al. Doughnut-shaped structure of a bacterial muramidase revealed by X-ray crystallography. Nature 367, 750–753 (1994).
Strickland, C. L.et al. Crystal structure of farnesyl protein transferase complexed with a CaaX peptide and farnesyl diphosphate analogue. Biochemistry 37, 16601–16611 (1998).
Peters, J. W., Stowell, M. H. & Rees, D. C. Aleucine-rich repeat variant with a novel repetitive protein structural motif. Nature Struct. Biol. 3, 991–994 (1996).
Huber, A. H., Nelson, W. J. & Weis, W. I. Three-dimensional structure of the armadillo repeat region of β-catenin. Cell 90, 871–882 (1997).
Conti, E., Uy, M., Leighton, L., Blobel, G. & Kuriyan, J. Crystallographic analysis of the recognition of a nuclear localization signal by the nuclear import factor karyopherin alpha. Cell 94, 193–204 (1998).
Groves, M. R.et al. The structure of the protein phosphatase 2A PR65/A subunit reveals the conformation of its 15 tandemly repeated HEAT motifs. Cell 96, 99–110 (1999).
Murzin, A. G., Brenner, S. E., Hubbard, T. & Chothia, C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J. Mol. Biol. 247, 536–540 (1995).
Das, A. K., Cohen, P. T. W. & Barford, D. The structure of the tetratricopeptide repeats of protein phosphatase 5: implications for TPR-mediated protein-protein interactions. EMBO J. 17, 1192–1199 (1998).
ter Haar, E., Musacchio, A., Harrison, S. C. & Kirchhausen, T. Atomic structure of clathrin: a β propeller terminal domain joins an α zigzag linker. Cell 95, 563–573 (1998).
Bucher, P., Karplus, K., Moeri, N. & Hofmann, K. Aflexible motif search technique based on generalized profiles. Comput. Chem. 20, 30–23 (1996).
Conibear, E. & Stevens, T. H. Multiple sorting pathways between the late Golgi and the vacuole in yeast. Biochim. Biophys. Acta 1404, 211–230 (1996).
Nakamura, N., Hirata, A., Ohsumi, Y. & Wada, Y. Vam2/Vps41p and Vam6/Vps39p are components of a protein complex on the vacuolar membranes and involved in the vacuolar assembly in the yeast Saccharomyces cerevisiae. J. Biol. Chem. 272, 11344–11349 (1997).
Winkler, F. K. & Stanley, K. K. Clathrin heavy chain, light chain interactions. EMBO J. 2, 1393–1400 (1983).
Kirchhausen, T.et al. Clathrin light chains LCa and LCb are similar, polymorphic, and share repeated heptad motifs. Science 236, 320–326 (1987).
Wilde, A.et al. EGF receptor signaling stimulates SRC kinase phosphorylation of clathrin, influencing clathrin redistribution and EGF uptake. Cell 96, 677–687 (1999).
Van Duyne, G. D., Standaert, R. F., Karplus, P. A., Schreiber, S. L. & Clardy, J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229, 105–124 (1993).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Terwilliger, T. C. Multiwavelength anomalous diffraction phasing of macromolecular structures: analysis of MAD data as single isomorphous replacement with anomalous scattering data using the MADMRG program. Methods Enzymol. 276, 530–537 (1997).
Brünger, A. T.et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Cryst. D 54, 905–921 (1998).
Kleywegt, G. J. & Jones, T. A. Asuper position. ESF/CCP4 Newsletter 31, 9–14 (1994).
Esnouf, M. An extensively modified version of MOLSCRIPT that includes greatly enhanced coloring capabilities. J. Mol. Graph. Model 15, 132–134 (1997).
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Merritt, E. A. & Murphy, M. E. P. RASTER3D version 2.0—a program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 (1994).
Acknowledgements
We thank F. Masiarz for mass spectrometry analysis; L.-W. Hung for assistance with data collection at beamline 5.0.2 of the Macromolecular Cyrstallography Facility at the Advanced Light Source (ALS is funded by the U.S. Department of Energy Office of Basic Energy Sciences); H. Bellamy and W. Weis for discussions on crystallographic data collection and analysis; T. Terwilliger and C. Weekes for correspondence on SOLVE and SnB2, respectively; M. Butte, for software assistance and comments on manuscript. This work was supported by the NIH (F.M.B., R.J.F.)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ybe, J., Brodsky, F., Hofmann, K. et al. Clathrin self-assembly is mediated by a tandemly repeated superhelix. Nature 399, 371–375 (1999). https://doi.org/10.1038/20708
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/20708
This article is cited by
-
Cryo-EM of multiple cage architectures reveals a universal mode of clathrin self-assembly
Nature Structural & Molecular Biology (2019)
-
Sequence evidence for common ancestry of eukaryotic endomembrane coatomers
Scientific Reports (2016)
-
Structure of clathrin coat with bound Hsc70 and auxilin: mechanism of Hsc70-facilitated disassembly
The EMBO Journal (2010)
-
Monomeric but not trimeric clathrin heavy chain regulates p53-mediated transcription
Oncogene (2008)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.