Abstract
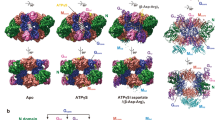
Porphobilinogen synthase (PBGS) catalyzes the first common step in the biosynthesis of tetrapyrroles (such as heme and chlorophyll). Although the predominant oligomeric form of this enzyme, as inferred from many crystal structures, is that of a homo-octamer, a rare human PBGS allele, F12L, reveals the presence of a hexameric form. Rearrangement of an N-terminal arm is responsible for this oligomeric switch, which results in profound changes in kinetic behavior. The structural transition between octamer and hexamer must proceed through an unparalleled equilibrium containing two different dimer structures. The allosteric magnesium, present in most PBGS, has a binding site in the octamer but not in the hexamer. The unprecedented structural rearrangement reported here relates to the allosteric regulation of PBGS and suggests that alternative PBGS oligomers may function in a magnesium-dependent regulation of tetrapyrrole biosynthesis in plants and some bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Battersby, A.R. Tetrapyrroles: the pigments of life. Nat. Prod. Rep. 17, 507–526 (2000).
Jaffe, E.K. The porphobilinogen synthase family of metalloenzymes. Acta Crystallogr. D 56, 115–128 (2000).
Akagi, R., Yasui, Y., Harper, P. & Sassa, S. A novel mutation of delta-aminolaevulinate dehydratase in a healthy child with 12% erythrocyte enzyme activity. Br. J. Haematol. 106, 931–937 (1999).
Schulze, A., Frommhold, D., Hoffman, G.F. & Mayatepek, E. Spectrophotometric microassay for delta-aminolevlinate dehydratase in dried-blood spots as confirmation for hereditary tyrosinemia type I. Clin. Chem. 47, 1424–1429 (2001).
Maruno, M. et al. Highly heterogeneous nature of delta-aminolevulinate dehydratase (ALAD) deficiencies in ALAD porphyria. Blood 97, 2972–2978 (2001).
Jaffe, E.K., Martins, J., Li, J., Kervinen, J. & Dunbrack, R.L. Jr. The molecular mechanism of lead inhibition of human porphobilinogen synthase. J. Biol. Chem. 276, 1531–1537 (2001).
Jaffe, E.K. et al. An artificial gene for human porphobilinogen synthase allows comparison of an allelic variation implicated in susceptibility to lead poisoning. J. Biol. Chem. 275, 2619–2626 (2000).
Jaffe, E.K. et al. Characterization of the role of the stimulatory magnesium of Escherichia coli porphobilinogen synthase. Biochemistry 34, 244–251 (1995).
Petrovich, R.M., Litwin, S. & Jaffe, E.K. Bradyrhizobium japonicum porphobilinogen synthase uses two Mg(II) and monovalent cations. J. Biol. Chem. 271, 8692–8699 (1996).
Frankenberg, N., Jahn, D. & Jaffe, E.K. Pseudomonas aeruginosa contains a novel type V porphobilinogen synthase with no required catalytic metal ions. Biochemistry 38, 13976–13982 (1999).
Erskine, P.T. et al. X-ray structure of 5-aminolaevulinate dehydratase, a hybrid aldolase. Nat. Struct. Biol. 4, 1025–1031 (1997).
Erskine, P.T. et al. X-ray structure of 5-aminolevulinic acid dehydratase from Escherichia coli complexed with the inhibitor levulinic acid at 2.0 Å resolution. Biochemistry 38, 4266–4276 (1999).
Erskine, P.T. et al. The Schiff base complex of yeast 5-aminolaevulinic acid dehydratase with laevulinic acid. Protein Sci. 8, 1250–1256 (1999).
Frankenberg, N. et al. High resolution crystal structure of a Mg2+-dependent porphobilinogen synthase. J. Mol. Biol. 289, 591–602 (1999).
Erskine, P.T. et al. MAD analyses of yeast 5-aminolaevulinate dehydratase: their use in structure determination and in defining the metal-binding sites. Acta Crystallogr. D 56, 421–430 (2000).
Kervinen, J. et al. Mechanistic basis for suicide inactivation of porphobilinogen synthase by 4,7-dioxosebacic acid, an inhibitor that shows dramatic species selectivity. Biochemistry 40, 8227–8236 (2001).
Erskine, P.T. et al. The X-ray structure of yeast 5-aminolaevulinic acid dehydratase complexed with two diacid inhibitors. FEBS Lett. 503, 196–200 (2001).
Erskine, P.T. et al. The X-ray structure of yeast 5-aminolaevulinic acid dehydratase complexed with substrate and three inhibitors. J. Mol. Biol. 312, 133–141 (2001).
Jaffe, E.K. et al. Species-specific inhibition of porphobilinogen synthase by 4-oxosebacic acid. J. Biol. Chem. 277, 19792–19799 (2002).
Frere, F. et al. Structure of porphobilinogen synthase from Pseudomonas aeruginosa in complex with 5-fluorolevulinic acid suggests a double Schiff base mechanism. J. Mol. Biol. 320, 327–247 (2002).
Murzin, A.G., Brenner, S.E., Hubbard, T. & Chothia, C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J. Mol. Biol. 247, 536–540 (1995).
Kervinen, J. et al. Porphobilinogen synthase from pea: expression from an artificial gene, kinetic characterization, and novel implications for subunit interactions. Biochemistry 39, 9018–9029 (2000).
Segel, I.H. Enzyme Kinetics 64–71 (Wiley, New York,1975).
Jaffe, E.K. An unusual phylogenetic variation in the metal ion binding sites of porphobilinogen synthase. Chem. Biol. 10, 25–34 (2003).
Frankenberg, N., Heinz, D.W. & Jahn, D. Production, purification, and characterization of a Mg2+-responsive porphobilinogen synthase from Psuedomonas aeruginosa. Biochemistry 38, 13968–13975 (1999).
Schneider, H.A. Enzymic capacities for chlorophyll biosynthesis. Activation and de novo synthesis of enzymes. Z. Naturforsch. 31, 55–63 (1976).
Papenbrock, J., Mock, H.P., Tanaka, R., Kruse, E. & Grimm, B. Role of magnesium chelatase activity in the early steps of the tetrapyrrole biosynthetic pathway. Plant Physiol. 122, 1161–1169 (2000).
Papenbrock, J. & Grimm, B. Regulatory network of tetrapyrrole biosynthesis—studies of intracellular signalling involved in metabolic and developmental control of plastids. Planta 213, 667–681 (2001).
Walker, D.A. Regulatory mechanisms in photosynthetic carbon metabolism. Curr. Top. Cell Regul. 11, 203–241 (1976).
Stolz, M. & Dornemann, D. Purification, metal cofactor, N-terminal sequence and subunit composition of a 5-aminolevulinic acid dehydratase from the unicellular green alga Scenedesmus obliquus, mutant C-2A′. Eur. J. Biochem. 236, 600–608 (1996).
Tamai, H., Shioi, Y. & Sasa, T. Purification and characterization of δ-aminolevulinic acid dehydratase from Chlorella regularis. Plant Cell Physiol. 20, 435–444 (1979).
Wu, J.W. et al. Crystal structure of a phosphorylated Smad2. Recognition of phosphoserine by the MH2 domain and insights on Smad function in TGF-β signaling. Mol. Cell 8,1277–1289 (2001).
Laue, T., Shaw, B.D., Ridgeway, T.M. & Pelletier, S.L. in Analytical Ultracentrifugation in Biochemistry and Polymer Science (eds. Harding, S.E., Rowe, A. & Horton, J.C.) 90–125 (Royal Society of Chemistry, Cambridge, UK, 1992).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Navaza, J. An automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Jones, T.A., Zou, J.Y., Cowan, S. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Acknowledgements
We thank J. Tannir (Fox Chase Cancer Center (FCCC)), S. Seeholzer (FCCC), T. Guszczynski (US National Cancer Institute (NCI)) and T. Copland (NCI) for technical support. The FCCC DNA synthesis facility, DNA sequencing facility and Research Secretarial Services were used in the preparation of this manuscript. This work was supported by the US National Institute of Environmental Health Sciences, the NCI, and by an appropriation from the Commonwealth of Pennsylvania.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Breinig, S., Kervinen, J., Stith, L. et al. Control of tetrapyrrole biosynthesis by alternate quaternary forms of porphobilinogen synthase. Nat Struct Mol Biol 10, 757–763 (2003). https://doi.org/10.1038/nsb963
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb963
This article is cited by
-
Heme biosynthesis depends on previously unrecognized acquisition of iron-sulfur cofactors in human amino-levulinic acid dehydratase
Nature Communications (2020)
-
Forkhead containing transcription factor Albino controls tetrapyrrole-based body pigmentation in planarian
Cell Discovery (2016)
-
Redox and metal-regulated oligomeric state for human porphobilinogen synthase activation
Amino Acids (2011)