Abstract
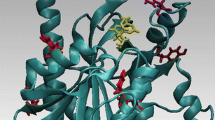
In order to gain insight into the light-driven repair of DNA by the enzyme DNA photolyase, the conformation of the photoactive cofactor FAD, a flavin adenine dinucleotide, has been studied by molecular dynamic simulations. In contrast to FAD in the gas phase and in water where the MD procedure yields various "open" I-shaped as well as "closed" U-shaped conformations, the calculations of FAD binding to the enzyme show essentially a single U-shaped conformation of this cofactor which, so far, is unique among FAD-carrying proteins. It is characteristic for this U-shaped conformation that the FAD components occupy opposite sides of the pocket in the surface of the protein which provides the binding site for the defect pyrimidine dimer structure on DNA. In fact, the calculated U-shaped conformation is very close to the one revealed by the X-ray structure analysis of DNA photolyase. Moreover, the simulations yield details on the binding of the photoactive isoalloxazine moiety and the dynamics of the amino acids forming the binding cavity of the enzyme.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 10 December 1997 / Accepted: 3 February 1998 / Published: 16 February 1998
Rights and permissions
About this article
Cite this article
Hahn, J., Michel-Beyerle, ME. & Rösch, N. Conformation of the Flavin Adenine Dinucleotide Cofactor FAD in DNA-Photolyase: A Molecular Dynamics Study. J Mol Med 4, 73–82 (1998). https://doi.org/10.1007/s008940050133
Issue Date:
DOI: https://doi.org/10.1007/s008940050133