Abstract
Molecular dynamics (MD) simulations on the complexes of glucoamylase II (471) from Aspergillus awamori var. X100 with two powerful inhibitors, 1-deoxynojirimycin and (+)-lentiginosine, have been performed, in order to build a model for these complexes in solution and to clarify the structure-activity relationship.
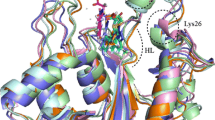
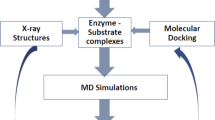
MD calculations were carried out for 105 ps, over a 15 Å sphere centered on the inhibitors. A 8 Å residue-based cut-off was used, and the calculations were performed with explicit inclusion of solvent molecules. The MD structure of the complex 1-deoxynojirimycin-glucoamylase shows only minor deviations from the available X-ray structure. The MD structure of the complex of (+)-lentiginosine-glucoamylase, obtained by docking the inhibitor into the active site, suggests us a suitable orientation for the molecule into the enzyme cavity, which can rationalize the high inhibition activity found for (+)-lentiginosine towards amyloglucosidase from A. niger.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 28 February 1997 / Accepted: 7 July 1997 / Published: 17 July 1997
Rights and permissions
About this article
Cite this article
Cardona, F., Goti, A., Brandi, A. et al. Molecular Dynamics Simulations on the Complexes of Glucoamylase II (471) from Aspergillus awamori var. X100 with 1-Deoxynojirimycin and Lentiginosine. J Mol Med 3, 249–260 (1997). https://doi.org/10.1007/s008940050037
Issue Date:
DOI: https://doi.org/10.1007/s008940050037