Summary
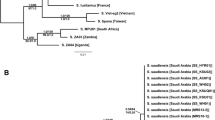
A fragment of the internal transcribed spacer (ITS-1) adjacent to the 5.8S rRNA gene of 20 myrmicine ant species was sequenced. Sequence comparisons were carried out between 11 species of the tribe Leptothoracini, five species of the tribe Tetramoriini, three species of the tribe Solenopsidini and one species of the tribe Myrmicini. Additionally, the formicine antCamponotus ligniperda (tribe Camponotini) was analyzed as an outgroup species. Among all investigated species, the fragment had a variable length of ≈ 230–380 bp with only a few conserved sequence elements. The sequences of this fragment were perfectly identical within four palearctic populations ofLeptothorax acervorum indicating that intraspecific variation is rather low. Within the species of Tetramoriini (includingAnergates atratulus) 94.1% of sequence positions were identical, 95.6% within the species of theLeptothorax s.str.-group and 64.6% within the species of theMyrafant-group. Phylogenetic analysis reveals that the social parasitesHarpagoxenus sublaevis, Doronomyrmex goesswaldi, D. kutteri andD. pacis, Chalepoxenus muellerianus as well asStrongylognathus alpinus andTeleutomyrmex schneideri are most closely related to the groups of their respective host species, which generally confirms the taxonomical classifications of the subfamily Myrmicinae based on morphological criteria. The taxonomical positions of the speciesA. atratulus has as yet been uncertain, however, sequence comparison of the ITS-1 fragment leads to the conclusion thatA. atratulus rather belongs to the tribe Tetramoriini than to the Solenopsidini.
Similar content being viewed by others
References
Arnheim, N., 1983. Concerted evolution of multigene families. In:Evolution of Genes and Proteins (M. Nei and R. K. Koehn, eds.), Sinauer, Sunderland, Mass. pp. 38–61.
Arnold, M. L., D. D. Shaw and M. Contreras, 1987. Ribosomal RNA-encoding DNA introgression across a narrow hybrid zone between two subspecies of grasshopper.Proc. Natl. Acad. Sci. USA 84:3946–3950.
Baur, A., A. Buschinger and F. K. Zimmermann, 1993. Molecular cloning and sequencing of 18S rDNA gene fragments from six different ant species.Ins. Soc. 40:325–335.
Bolton, B., 1976. The ant tribe Tetramoriini (Hymenoptera: Formicidae): constituent genera, review of smaller genera and revision ofTriglyphothrix FOREL.Bull. Brit. Mus. nat. Hist. (Ent.) 34:281–379.
Buschinger, A., 1987. Biological arguments for a systematic rearrangement of the ant tribe Leptothoracini. In:Chemistry and Biology of Social Insects (J. Eder and H. Rembold, eds.), J. Peperny, München, p. 43.
Buschinger, A., 1990. Sympatric speciation and radiative evolution of socially parasitic ants — Heretic hypotheses and their factual background.Z. Zool. Syst. Evolut.-forsch. 28:241–260.
Coen, E. S., T. Strachan and G. A. Dover, 1982. Dynamics of concerted evolution of ribosomal DNA and histone gene families in themelanogaster species subgroup ofDrosophila.J. Mol. Biol. 158:17–35.
Devereux, J., P. Haeberli and O. Smithies, 1984. A comprehensive set of sequence analysis programmes for the VAX.Nucleic Acid Res. 12:387–395.
Douwes, P. and B. Stille, 1987. The use of enzyme electrophoresis inLeptothorax classification. In:Chemistry and Biology of Social Insects (J. Eder and H. Rembold, eds.), J. Peperny, München, pp. 29–30.
Dowling, T. E., C. Moritz and J. Palmer, 1990. Nucleic acids II: Restriction site analysis. In:Molecular Systematics (D. M. Hillis and C. Moritz, eds.), Sinauer, Sunderland, Mass. pp. 250–317.
Dover, G. and E. Coen, 1981. Springcleaning ribosomal DNA. A model for multigene evolution?Nature 290:731–732.
Emery, C., 1909. Über den Ursprung der dulotischen, parasitischen und myrmekophilen Ameisen.Biol Zbl. 29:352–362.
Emery, C., 1921-1922. Hymenoptera, fam. Formicidae, subfam. Myrmicinae. In:Genera Insec-torum, no. 174 (P. Wytsman, ed.), Louis Desmet-Verteneul, Brussels. 397 pp.
Ettershank, G., 1966. A generic revision of the world Myrmicinae related toSolenopsis andPheidologeton.Aust. J. Zool. 14:73–171.
Felsenstein, J., 1985. Confidence limits on phylogenies: an approach using the bootstrap.Evolution 39:783–791.
Felsenstein, J., 1989. PHYLIP — Phylogeny Inference Package (Version 3.2).Cladistics 5:164–166.
Felsenstein, J., 1993. PHYLIP (Phylogeny Inference Package) Version 3.5 c. Dep. of Genetics, University of Washington, Seattle.
Fitch, W. M. and E. Margoliash, 1967. Construction of phylogenetic trees.Science 155:279–284.
Fujiwara, H., Y. Kawata and H. Ishikawa, 1982. Primary structure of the 5.8S rRNA from the silk gland ofBombyx mori.Nucleic Acid Res. 10:2415–2418.
Gonzales, I. L., C. Chambers, J. L. Gorski, D. Stambolian, R. D. Schmickel and J. E. Sylvester, 1990. Sequence and structure correlation of human ribosomal transcribed spacers.J. Mol. Biol. 212:27–35.
Heinze, J., 1991. Biochemical studies on the relationship between socially parasitic ants and their hosts.Biochem. Syst. Ecol. 19:195–206.
Hillis, D. M. and M. T. Dixon, 1991. Ribosomal DNA: Molecular evolution and phylogenetic inference.Quart. Rev. Biol. 66:411–453.
Hölldobler, B. and E. O. Wilson, 1990.The Ants. The Belknap Press of Harvard University Press, Cambridge, Mass. 732 pp.
Jukes, T. H. and R. C. Cantor, 1969. Evolution of protein molecules. In:Mammalian protein metabolism (H. N. Munoro, ed.), Academic Press, New York. pp. 21–132.
Kuperus, W. R. and W. Chapco, 1994. Usefulness of internal transcribed spacer regions of ribosomal DNA in melanopline (Orthoptera: Acrididae) systematics.Ann. Entomol. Soc. Am. 87:751–754.
Kutter, H., 1969. Die sozialparasitischen Ameisen der Schweiz.Neujahrsbl. Naturf. Ges. Zürich 1969:1–62.
Kutter, H., 1977. Hymenoptera: Formicidae. In:Insecta Helvetica, vol. 6 (W. Sauter, ed.), Schweiz. Ent. Ges., Zürich. 289 pp.
Kwok, S. and R. Higuchi, 1989. Avoiding false positives with PCR.Nature 339:237–238.
Mullis, K. B. and F. Faloona, 1987. Specific synthesis of DNA in vitro via a polymerase-catalyzed chain reaction.Methods Enzymol. 155:335–350.
Muster, W., K. Boon, A. F. M. van der Sande, H. van Heerikuizen and R. J. Planta, 1990. Functional analysis of transcribed spacers of yeast ribosomal DNA.EMBO J. 9:3989–3996.
Nazar, R. N., T. O. Sitz and H. Busch, 1976. Sequence homologies in the mammalian 5.8S ribosomal RNA.Biochemistry 15:505–508.
Pavlakis, G. N., B. R. Jourdan, R. M. Wurst and C. Vournakis, 1979. Sequence and secondary structure ofDrosophila melanogaster 5.8S and 28S rRNAs and of the processing site between them.Nucleic Acid Res. 7:2213–2238.
Pearson, W. R. and D. J. Lipman, 1988. Improved tools for biological sequence comparison.Proc. Natl. Sci. USA 85:2444–2448.
Ritland, C. E., K. Ritland and N. A. Straus, 1993. Variation in the ribosomal internal transcribed spacers (ITS1 and ITS2) among eight taxa of theMimulus guttatus species complex.Mol Biol. Evol. 10:1273–1288.
Rubin, G. M., 1973. The nucleotide sequence ofSaccharomyces cerevisiae 5.8S ribosomal nucleic acid.J. Biol. Chem. 248:3860–3875.
Saiki, R. K., S. Scharf, F. Faloona, K. B. Mullis, G. T. Horu, H. A. Erlich and N. Arnheim, 1985. Enzymatic amplification of β-globulin sequences and restriction site analysis for diagnosis of sickle cell anemia.Science 230:1350–1354.
Saitou, N. and M. Nei, 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees.Mol. Biol. Evol. 4:406–425.
Sanetra, M., J. Heinze and A. Buschinger, 1994. Enzyme polymorphism in the ant genusTetramorium Mayr and its social parasites.Biochem. Syst. Ecol. 22:753–759.
Sambrook, J., E. F Fritsch and T. Maniatis, 1989.Molecular cloning — a Laboratory Manual, Second Edition. Cold Spring Harbour Laboratory Press.
Sanger, F., S. Nicklen and M. Coulson, 1977. DNA sequencing with chain terminating inhibitors.Proc. Natl. Acad. Sci. USA 162:5463–5467.
Smith, T. F. and M. S. Waterman, 1981. Comparison of bio-sequences.Adv. Appl. Math. 2:482–488.
Tautz, D., C. Tautz, D. Webb and G. A. Dover, 1987. Evolutionary divergence of promoters and spacers in the rDNA family of fourDrosophila species. Implications for molecular coevolution in multigene families.J. Mol. Biol. 195:525–542.
Wesson, D. M., D. K. McLain, J. H. Oliver, J. Piesman and F. H. Collins, 1993. Investigation of the validity of species status ofIxodes dammini (Acari: Ixodidae) using rDNA.Proc. Natl. Acad. Sci. USA 90:10221–10225.
Wesson, D. M., C. H. Porter and F. H. Collins, 1992. Sequence and secondary structure comparison of the ITS rDNA in Mosquitoes (Diptera: Culicidae).Mol. Phylogen. Evol. 1:253–269.
Wheeler, G. C. and J. Wheeler, 1985. A simplified conspectus of the Formicidae.Trans. Am. Entomol. Soc. 111:255–264.
Author information
Authors and Affiliations
Additional information
Author who provided the sequence data from his doctoral thesis.
Rights and permissions
About this article
Cite this article
Baur, A., Sanetra, M., Chalwatzis, N. et al. Sequence comparisons of the internal transcribed spacer region of ribosomal genes support close relationships between parasitic ants and their respective host species (Hymenoptera: Formicidae). Ins. Soc 43, 53–67 (1996). https://doi.org/10.1007/BF01253956
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01253956