Abstract
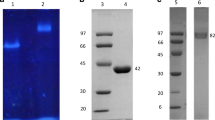
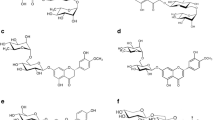
Bacteria represent an underexplored source of diglycosidases. Twenty-five bacterial strains from the genera Actinoplanes, Bacillus, Corynebacterium, Microbacterium, and Streptomyces were selected for their ability to grow in diglycosylated flavonoids-based media. The strains Actinoplanes missouriensis and Actinoplanes liguriae exhibited hesperidin deglycosylation activity (6-O-α-L-rhamnosyl-β-D-glucosidase activity, EC 3.2.1.168), which was 3 to 4 orders of magnitude higher than the corresponding monoglycosidase activities. The diglycosidase production was confirmed in A. missouriensis by zymographic assays and NMR analysis of the released disaccharide, rutinose. The gene encoding the 6-O-α-L-rhamnosyl-β-D-glucosidase was identified in the genome sequence of A. missouriensis 431T (GenBank accession number BAL86042.1) and functionally expressed in Escherichia coli. The recombinant protein hydrolyzed hesperidin and hesperidin methylchalcone, but not rutin, which indicates its specificity for 7-O-rutinosylated flavonoids. The protein was classified into the glycoside hydrolase family 55 (GH55) in contrast to the known eukaryotic diglycosidases, which belong to GH1 and GH5. These findings demonstrate that organisms other than plants and filamentous fungi can contribute to an expansion of the diglycosidase toolbox.





Similar content being viewed by others
References
Bianchetti CM, Takasuka TE, Deutsch S, Udell HS, Yik EJ, Bergeman LF, Fox BG (2015) Active site and laminarin binding in glycoside hydrolase family 55. J Biol Chem 290:11819–11832. doi:10.1074/jbc.M114.623579
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: John M. Walker (ed): The Proteomics Protocols Handbook, Humana Press, pp 571–607
González C, Martínez A, Vázquez F, Baigori M, Figueroa LIC (1996) New method of screening and differentiation of exoenzymes from industrial strains. Biotechnol Tech 10:519–522. doi:10.1007/BF00159517
Hemingway KM, Alston MJ, Chappell CG, Taylor AJ (1999) Carbohydrate-flavour conjugates in wine. Carbohydr Polymer 38:283–286. doi:10.1016/S0144-8617(98)00103-9
Ishida T, Fushinobu S, Kawai R, Kitaoka M, Igarashi K, Samejima M (2009) Crystal structure of glycoside hydrolase family 55 β-1,3-glucanase from the basidiomycete Phanerochaete chrysosporium. J Biol Chem 284:10100–10109. doi:10.1074/jbc.M808122200
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. doi:10.1038/227680a0
Lombard V, Golaconda RH, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:D490–D495
Ma SJ, Mizutani M, Hiratake J, Hayashi K, Yagi K, Watanabe N, Sakata K (2001) Substrates specificity of β-primeverosidase, a key enzyme in aroma formation during oolong tea and black tea manufacturing. Biosci Biotechnol Biochem 65:2719–2729
Manthey JA, Grohmann K (1996) Concentrations of hesperidin and other orange peel flavonoids in citrus processing by products. J Agric Food Chem 44:811–814. doi:10.1021/jf950572g
Mazzaferro L, Piñuel L, Minig M, Breccia JD (2010) Extracellular monoenzyme deglycosylation system of 7-O-linked flavonoid β-rutinosides and its disaccharide transglycosylation activity from Stilbella fimetaria. Arch Microbiol 2010;192:383–393 Erratum in: Arch Microbiol (2011) 193:461. doi:10.1007/s00203-010-0567-7
Mazzaferro L, Breccia JD (2011) Functional and biotechnological insights into diglycosidases. Biocat Biotrans 29:103–112. doi:10.3109/10242422.2011.594882
Mazzaferro L, Piñuel L, Erra-Balsells R, Giudicessi SL, Breccia JD (2012) Transglycosylation specificity of Acremonium sp. α-rhamnosyl-β-glucosidase and its application to the synthesis of the new fluorogenic substrate 4-methylumbelliferyl-rutinoside. Carbohydr Res 347:69–75. doi:10.1016/j.carres.2011.11.008
Merkens H, Kappl R, Jakob RP, Schmid FX, Fetzner S (2008) Quercetinase QueD of Streptomyces sp. FLA, a monocupin dioxygenase with a preference for nickel and cobalt. Biochemistry 47:12185–12196. doi:10.1021/bi801398x
Minig M, Walker D, Ledesma P, Martínez MA, Breccia JD (2009) Bacterial isolates from Ethiopian soda lake producers of alkaline-active β-glucanases resistant to chelating and surfactant compounds. Res J Microbiol 4:194–201. doi:10.3923/jm.2009.194.201
Minig M, Mazzaferro LS, Erra-Balsells R, Petroselli G, Breccia JD (2011) α-Rhamnosyl-β-glucosidase-catalyzed reactions for analysis and biotransformations of plant-based foods. J Agr Food Chem 59:11238–11243
Mizutani M, Nakanishi H, Ema J, Ma SJ, Noguchi E, Inohara-Ochiai M, Fukuchi-Mizutani M, Nakao M, Sakata K (2002) Cloning of β-primeverosidase from tea leaves, a key enzyme in tea aroma formation. Plant Physiol 130:2164–2176
Nam HK, Hong SY, Shin KC, Oh DK (2012) Quercetin production from rutin by a thermostable β-rutinosidase from Pyrococcus furiosus. Biotechnol Lett 34:483–489. doi:10.1007/s10529-011-0786-2
Narikawa T, Shinoyama H, Fujii T (2000) A β-rutinosidase from Penicillium rugulosum IFO 7242 that is a peculiar flavonoid glycosidase. Biosci Biotechnol Biochem 64:1317–1319. doi:10.1271/bbb.64.1317
Okazaki K, Nishimura N, Matsuoka F, Hayakawa S (2007) Cloning and characterization of the gene encoding endo-β-1,3-glucanase from Arthrobacter sp. NHB-10. Biosci Biotechnol Biochem 71(6):1568–1571. doi:10.1271/bbb.70030
Orrillo AG, Ledesma P, Delgado OD, Spagna G, Breccia JD (2007) Cold-active α-L-rhamnosidase from psychrotolerant bacteria isolated from a sub-Antarctic ecosystem. Enz Microb Technol 40:236–241. doi:10.1016/j.enzmictec.2006.04.002
Petersen TN, Brunak S, Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786. doi:10.1038/nmeth.1701
Rao JR, Cooper JE (1994) Rhizobia catabolize nod gene-inducing flavonoids via C-ring fission mechanisms. J Bacteriol 176:5409–5413. doi:0021–9193/94/$04.00+0
Rose K, Fetzner S (2006) Identification of linear plasmid pAM1 in the flavonoid degrading strain Actinoplanes missouriensis (DSM 43046). Plasmid 55:249–254. doi:10.1016/j.plasmid.2005.10.003
Sambrook J, Russell D (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Sang-Joon L, Omorl T, Kodama T (1990) Purification and some properties of rutinosidase from Arthrobacter sp. Kor J Appl Microbiol Biotech 18:360–367
Sarry JE, Gunata Z (2004) Plant and microbial glycoside hydrolases: volatile release from glycosidic aroma precursors. Food Chem 87:509–521. doi:10.1016/j.foodchem.2004.01.003
Schubert M, Melnikova AN, Mesecke N, Zubkova EK, Fortte R, Batashev DR, Barth I, Sauer N, Gamalei YV, Mamushina NS, Tietze LF, Voitsekhovskaja OV, Pawlowski K (2010) Two novel disaccharides, rutinose and methylrutinose, are involved in carbon metabolism in Datisca glomerata. Planta 231:507–521. doi:10.1007/s00425-009-1085-1
Šimčíková D, Kotik M, Weignerová L, Halada P, Pelantová H, Adamcová K, Křen V (2015) α-L-Rhamnosyl-β-D-glucosidase (rutinosidase) from Aspergillus niger: characterization and synthetic potential of a novel diglycosidase. Adv Synth Catal 357(1):107–117. doi:10.1002/adsc.201400566
Wang D, Kurasawa E, Yamaguchi Y, Kubota K, Kobayashi A (2001) Analysis of glycosidically bound aroma precursors in tea leaves. 2. Changes in glycoside contents and glycosidase activities in tea leaves during the black tea manufacturing process. J Agric Food Chem 49:1900–1903. doi:10.1021/jf001077+
Westlake DWS, Talbot G, Blakley ER, Simpson FJ (1959) Microbial decomposition of rutin. Can J Microbiol 5:621–629. doi:10.1139/m59-076
Yamamoto S, Okada M, Usui T, Sakata K, Toumoto A, Tsuruhami K (2006) Diglycosidase isolated from microorganisms. United States Patent 71090142006 Sep 19.
Yamamura H, Ohnishi Y, Ishikawa J, Ichikawa N, Ikeda H, Sekine M, Harada T, Horinouchi S, Otoguro M, Tamura T, Suzuki K, Hoshino Y, Arisawa A, Nakagawa Y, Fujita N, Hayakawa M (2012) Complete genome sequence of the motile actinomycete Actinoplanes missouriensis 431T (= NBRC 102363T). SIGS 7:294–303. doi:10.4056/sigs.3196539
Acknowledgments
This work was supported by the National University of La Pampa (UNLPam), the National Council of Scientific and Technical Research (CONICET), and The National Agency for Science and Technology Promotion (ANPCyT) of Argentina. Funding of the bilateral cooperation project 7AMB13AR005 (ARC/12/04) by the Ministry of Education, Youth and Sports (MEYS) of the Czech Republic and the Ministry of Science, Technology and Innovation (MINCYT) of Argentina is acknowledged. This work was also supported by the MEYS projects LD13042 and LD15085. Further, the authors also thank the support for the networking by the European COST Chemistry project MultiGlycoNano CM1102.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no competing interests.
Electronic supplementary material
ESM 1
(PDF 546 kb)
Rights and permissions
About this article
Cite this article
Neher, B.D., Mazzaferro, L.S., Kotik, M. et al. Bacteria as source of diglycosidase activity: Actinoplanes missouriensis produces 6-O-α-l-rhamnosyl-β-d-glucosidase active on flavonoids. Appl Microbiol Biotechnol 100, 3061–3070 (2016). https://doi.org/10.1007/s00253-015-7088-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-7088-x