Abstract
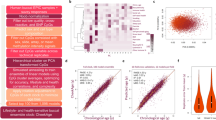
Monitoring health conditions is essential to detect early asymptomatic stages of a disease. To achieve this, blood, urine and breath samples are commonly used as a routine clinical diagnostic. These samples offer the opportunity to detect specific metabolites related to diseases and provide a better understanding of their development. Although blood samples are commonly used routinely to monitor health, the implementation of a relatively noninvasive technique, such as exhaled breath condensate (EBC) analysis, may further benefit the well-being of both humans and other animals. EBC analysis can be used to track possible physical or biochemical alterations caused by common diseases of the bottlenose dolphin (Tursiops truncatus), such as infections or inflammatory-mediated processes. We have used an untargeted metabolomic method with liquid chromatography–mass spectrometry analysis of EBC samples to determine biomarkers related to disease development. In this study, five dolphins under human care were followed up for 1 year. We collected paired blood, physical examination information, and EBC samples. We then statistically correlated this information to predict specific health alterations. Three dolphins provided promising case study information about biomarkers related to cutaneous infections, respiratory infections, dental disease, or hormonal changes (pregnancy). The use of complementary liquid chromatography platforms, with hydrophilic interaction chromatography and reverse-phased columns, allowed us to detect a wide spectrum of EBC biomarker compounds that could be related to these health alterations. Moreover, these two analytical techniques not only provided complementary metabolite information but in both cases they also provided promising diagnostic information for these health conditions.

Collection of the exhaled condensed breath from a bottlenose dolphin from U.S. Navy Marine Mammal Program (MMP)






Similar content being viewed by others
References
Nagana Gowda GA, Zhang S, Gu H, Asiago V, Shanaiah N, Raftery D. Metabolomics-based methods for early disease diagnostics: a review. Expert Rev Mol Diagn. 2008;8(5):617–33. https://doi.org/10.1586/14737159.8.5.617.
Zhang A, Sun H, Yan G, Wang P, Wang X. Metabolomics for biomarker discovery: moving to the clinic. Biomed Res Int. 2015;354671. https://doi.org/10.1155/2015/354671.
Miller DL, Ewing RY, Bossart GD. Emerging and resurging diseases. In: Dierauf L, Gulland FMD, editors. CRC handbook of marine mammal medicine. Boca Raton: CRC Press; 2001. p. 15–30.
Johnson WR, Torralba M, Fair PA, Bossart GD, Nelson KE, Morris PJ. Novel diversity of bacterial communities associated with bottlenose dolphin upper respiratory tracts. Environ Microbiol Rep. 2009;1(6):555–62. https://doi.org/10.1111/j.1758-2229.2009.00080.x.
Bik EM, Costello EK, Switzer AD, Callahan BJ, Holmes SP, Wells RS, et al. Marine mammals harbor unique microbiotas shaped by and yet distinct from the sea. Nat Commun. 2016;7:10516. https://doi.org/10.1038/ncomms10516.
Pasamontes A, Aksenov AA, Schivo M, Rowles T, Smith CR, Schwacke LH, et al. Noninvasive respiratory metabolite analysis associated with clinical disease in cetaceans: a Deepwater Horizon oil spill study. Environ Sci Technol. 2017;51(10):5737–46. https://doi.org/10.1021/acs.est.6b06482.
Wild CP. Complementing the genome with an “exposome”: the outstanding challenge of environmental exposure measurement in molecular epidemiology. Cancer Epidemiol Biomark Prev. 2005;14(8):1847–50. https://doi.org/10.1158/1055-9965.epi-05-0456.
King D, Aldridge B, Kennedy-Stoskopf S, Stott JL. Immunology. In: Dierauf L, Gulland FMD, editors. CRC handbook of marine mammal medicine. Boca Raton: CRC Press; 2001. p. 237–52.
Lima N, Rogers T, Acevedo-Whitehouse K, Brown MV. Temporal stability and species specificity in bacteria associated with the bottlenose dolphins respiratory system. Environ Microbiol Rep. 2012;4(1):89–96. https://doi.org/10.1111/j.1758-2229.2011.00306.x.
Zamuruyev K, Aksenov A, Baird M, Pasamontes A, Parry C, Foutouhi S, et al. Enhanced non-invasive respiratory sampling from bottlenose dolphins for breath metabolomics measurements. J Breath Res. 2016;10(4):046005.
Cumeras R, Cheung W, Gulland F, Goley D, Davis C. Chemical analysis of whale breath volatiles: a case study for non-invasive field health diagnostics of marine mammals. Metabolites. 2014;4(3):790.
Hunt KE, Moore MJ, Rolland RM, Kellar NM, Hall AJ, Kershaw J, et al. Overcoming the challenges of studying conservation physiology in large whales: a review of available methods. Conserv Physiol. 2013;1(1):cot006-cot. https://doi.org/10.1093/conphys/cot006.
Richard JT, Robeck TR, Osborn SD, Naples L, McDermott A, LaForge R, et al. Testosterone and progesterone concentrations in blow samples are biologically relevant in belugas (Delphinapterus leucas). Gen Comp Endocrinol. 2017;246:183–93. https://doi.org/10.1016/j.ygcen.2016.12.006.
Wilson B, Arnold H, Bearzi G, Fortuna CM, Gaspar R, Ingram S, et al. Epidermal diseases in bottlenose dolphins: impacts of natural and anthropogenic factors. Proc R Soc B Biol Sci. 1999;266(1423):1077–83.
Venn-Watson SK, Jensen ED, Ridgway SH. Evaluation of population health among bottlenose dolphins (Tursiops truncatus) at the United States Navy Marine Mammal Program. J Am Vet Med Assoc. 2011;238(3):356–60. https://doi.org/10.2460/javma.238.3.356.
Schwacke LH, Smith CR, Townsend FI, Wells RS, Hart LB, Balmer BC, et al. Health of common bottlenose dolphins (Tursiops truncatus) in Barataria Bay, Louisiana, following the Deepwater Horizon oil spill. Environ Sci Technol. 2014;48(1):93–103. https://doi.org/10.1021/es403610f.
Mazzaro LM, Johnson SP, Fair PA, Bossart G, Carlin KP, Jensen ED, et al. Iron indices in bottlenose dolphins (Tursiops truncatus). Comp Med. 2012;62(6):508–15.
Goldstein JD, Schaefer AM, McCulloch SD, Fair PA, Bossart GD, Reif JS. Clinicopathologic findings from Atlantic bottlenose dolphins (Tursiops Truncatus) with cytologic evidence of gastric inflammation. J Zoo Wildl Med. 2012;43(4):730–8. https://doi.org/10.1638/2011-0054R.1.
Schivo M, Aksenov A, Yeates L, Pasamontes A, Davis C. Diabetes and the metabolic syndrome: possibilities of a new breath test in a dolphin model. Front Endocrinol. 2013;4:163. https://doi.org/10.3389/fendo.2013.00163.
Aksenov AA, Yeates L, Pasamontes A, Siebe C, Zrodnikov Y, Simmons J, et al. Metabolite content profiling of bottlenose dolphin exhaled breath. Anal Chem. 2014;86(21):10616–24. https://doi.org/10.1021/ac5024217.
Raverty SA, Rhodes LD, Zabek E, Eshghi A, Cameron CE, Hanson MB, et al. Respiratory microbiome of endangered southern resident killer whales and microbiota of surrounding sea surface microlayer in the eastern North Pacific. Sci Rep. 2017;7(1):394. https://doi.org/10.1038/s41598-017-00457-5.
Kubáň P, Foret F. Exhaled breath condensate: Determination of non-volatile compounds and their potential for clinical diagnosis and monitoring. A review. Anal Chim Acta. 2013;805:1–18. https://doi.org/10.1016/j.aca.2013.07.049.
Wehrens R, Hageman JA, van Eeuwijk F, Kooke R, Flood PJ, Wijnker E, et al. Improved batch correction in untargeted MS-based metabolomics. Metabolomics. 2016;12:88. https://doi.org/10.1007/s11306-016-1015-8.
Jackson JE. A user's guide to principal components. Hoboken: Wiley; 2004.
Bartel J, Krumsiek J, Theis FJ. Statistical methods for the analysis of high-throughput metabolomics data. Comput Struct Biotechnol J. 2013;4(5):e201301009. https://doi.org/10.5936/csbj.201301009.
Barker M, Rayens W. Partial least squares for discrimination. J Chemom. 2003;17(3):166–73. https://doi.org/10.1002/cem.785.
Gika HG, Theodoridis GA, Plumb RS, Wilson ID. Current practice of liquid chromatography–mass spectrometry in metabolomics and metabonomics. J Pharm Biomed Anal. 2014;87:12–25. https://doi.org/10.1016/j.jpba.2013.06.032.
Sweeney JC, Ridgway SH. Common diseases of small cetaceans. J Am Vet Med Assoc. 1975;167(7):533–40.
Nemeth E, Ganz T. Anemia of inflammation. Hematol Oncol Clin North Am. 2014;28(4):671–681, vi. https://doi.org/10.1016/j.hoc.2014.04.005.
Horváth I, Barnes PJ, Loukides S, Sterk PJ, Högman M, Olin A-C et al. A European Respiratory Society technical standard: exhaled biomarkers in lung disease. Eur Respir J. 2017;49(4). https://doi.org/10.1183/13993003.00965-2016.
Acknowledgements
Support for these investigations was provided by the Office of Naval Research grant N-00014-13-1-0580 (CED, SV-W, BCW), National Institutes of Health (NIH) grants 1U01EB022003-01, UL1RR024146-06, 1P30ES023513-01A1, and UG3-OD023365 (CED) and T32-HL007013 and T32-ES007059 (MS), and University of California, Davis School of Medicine and NIH 8KL2TR000134-07 K12 mentored training award and NIH grant 1K23HL127185-01A1 (MS). Student support was provided by NIH award T32 HL07013 (KOZ) and NIH award P42ES004699 (KOZ).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This work was performed under an animal care and use protocol reviewed and approved by the US Navy Marine Mammal Program Institutional Animal Care and Use Committee and the U.S. Navy Bureau of Medicine and Surgery. The US Navy has elected to file patent application 14/859,612 on the dolphin breath sampler used in this work. The patent rights are assigned to the government, and the authors declare they have no financial conflicts of interest.
Electronic supplementary material
ESM 1
(PDF 65 kb)
Rights and permissions
About this article
Cite this article
Borras, E., Aksenov, A.A., Baird, M. et al. Exhaled breath condensate methods adapted from human studies using longitudinal metabolomics for predicting early health alterations in dolphins. Anal Bioanal Chem 409, 6523–6536 (2017). https://doi.org/10.1007/s00216-017-0581-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-017-0581-6