Abstract
Sustainable canola production is essential to meet growing human demands for vegetable oil, biodiesel, and meal for stock feed markets. Blackleg, caused by the fungal pathogen, Leptosphaeria maculans is a devastating disease that can lead to significant yield loss in many canola production regions worldwide. Breakdown of race-specific resistance to L. maculans in commercial cultivars poses a constant threat to the canola industry. To identify new alleles, especially for quantitative resistance (QR), we analyzed 177 doubled haploid (DH) lines derived from an RP04/Ag-Outback cross. DH lines were evaluated for QR under field conditions in three experiments conducted at Wagga Wagga (2013, 2014) and Lake Green (2015), and under shade house conditions using the ‘ascospore shower’ test. DH lines were also characterized for qualitative R gene-mediated resistance via cotyledon tests with two differential single spore isolates, IBCN17 and IBCN76, under glasshouse conditions. Based on 18,851 DArTseq markers, a linkage map representing 2,019 unique marker bins was constructed and then utilized for QTL detection. Marker regression analysis identified 22 significant marker associations for resistance, allowing identification of two race-specific resistance R genes, Rlm3 and Rlm4, and 21 marker associations for QR loci. At least three SNP associations for QR were repeatedly detected on chromosomes A03, A07 and C04 across phenotyping environments. Physical mapping of markers linked with these consistent QR loci on the B. napus genome assembly revealed their localization in close proximity of the candidate genes of B. napus BnaA03g26760D (A03), BnaA07g20240D (A07) and BnaC04g02040D (C04). Annotation of these candidate genes revealed their association with protein kinase and jumonji proteins implicated in defense resistance. Both Rlm3 and Rlm4 genes identified in this DH population did not show any association with resistance loci detected under either field and/or shade house conditions (ascospore shower) suggesting that both genes are ineffective in conferring resistance to L. maculans in Australian field conditions. Taken together, our study identified sequence-based molecular markers for dissecting R and QR loci to L. maculans in a canola DH population from the RP04/Ag-Outback cross.
Similar content being viewed by others
Introduction
Blackleg, caused by the fungal pathogen Leptosphaeria maculans, is a major disease of the canola (Brassica napus L., AnAnCnCn, 2n = 4× = 38) industry especially in Australia, Canada, and France. Developing new cultivars resistant to blackleg is recognized as the most economically and environmentally safe strategy to control yield losses. Traditional breeding for blackleg resistance largely relies on the selection of elite ‘good looking’ lines in disease nurseries where there is high disease pressure from a diverse pathogen population. Utilizing this strategy, several canola varieties have been developed and released for commercial cultivation with ratings of moderate- to highly-resistant (to blackleg) without any prior knowledge of either race-specific resistance (qualitative resistance; R) and/or race non-specific resistance (quantitative resistance; QR) genes.
R gene mediated resistance generally follows the classical gene-to-gene hypothesis; although genetic interactions between R genes have been reported in B. napus - L. maculans pathosystem1,2. R genes often have high heritability due to the major allelic effects, which can be tested efficiently and accurately at the cotyledon stage3,4. When those resistant genotypes harboring R gene(s) are subsequently grown they also exhibit a resistance response at the adult plant stage under controlled conditions3,4,5. However, a consistent resistance response may not be detected when plants are grown under natural field conditions, as those individuals are constantly challenged with in-coming inoculum from sexually reproducing diverse races of L. maculans throughout the Australian canola growing season (March-November). Conversely, QR is usually controlled by several genes of smaller allelic effect however these QR alleles together confer more durable resistance at the adult plant stage under field conditions. Due to strong genotype × environment interactions, the effects of QR loci detected in the mapping populations are often relatively small compared to R genes3,5,6,7,8,9,10,11. Currently, deployment of QR is preferred by canola breeding programs due to its durability and lower chance of breakdown of resistance to the highly-diverse nature of L. maculans as observed in some Australian and French cultivars12,13,14.
Quantitative resistance has been genetically mapped in a relatively limited number of structured bi-parental doubled haploid (DH) populations of canola using QTL linkage mapping3,4,8,10, and by genome-wide association approaches5,15,16,17. These studies showed that (i) multiple QTL are located on all 19 chromosomes of B. napus which control resistance to L. maculans, and (ii) QTL allelic effects vary depending upon the environment and the method of evaluation (per cent survival or internal infection scores). In addition, due to the usage of different genetic populations with spring, semi-winter, or winter backgrounds plus a range of molecular marker systems, it has been difficult to compare the locations of QTL for QR in canola from different studies. Recently, the genome of B. napus cv. Darmor-bzh, along with the physical positions of 425 R genes implicated in disease resistance, has been published18,19. This valuable resource provides an opportunity to compare QTL regions involved in resistance to L. maculans across populations, and to identify putative candidate genes underlying R and QR loci.
In this study, we identified loci associated with R and QR using linkage analysis and located their physical map positions on the Darmor genome assembly enabling the identification of putative candidate genes. Comparisons were also made between the significant QTL detected in the present study with those from previous studies which utilized sequence-based markers.
Materials and methods
Plant materials and single spore isolates
A DH population (RADH) comprising 177 lines was developed from a F1 cross between two Australian genotypes RP04 and Ag-Outback using microspore culture. This breeding DH population was generated at the Agriculture Victoria, Grains Innovation Park, Horsham, Australia. RP04 is an advanced breeding line developed by Agriculture Victoria (Ag-Vic) while Ag-Outback is a commercial canola variety developed from the cross: AGA95-1/Monty by AgSeed Pty Ltd (now Nuseed Pty Ltd, Australia) and released for cultivation in Australia20. Both parental genotypes were moderately resistant to blackleg disease under field conditions (Salisbury, unpublished data) and also shown to differ in fractional ground cover21, flowering time, plant height and seed yield (Raman et al., unpublished data). Seed of DH lines was multiplied in caged tents to avoid any cross pollination. A set of 10 L. maculans differential isolates22 was procured from Marcroft Grains Pathology, Horsham and maintained on 10% V8 juice agar (Campbell, Australia) at the Wagga Wagga Agricultural Institute (WWAI) for phenotyping experiments.
Phenotypic evaluation of DH lines
Evaluation of RADH population for resistance under field conditions
The RADH population, along with its parental lines, RP04 and Ag-Outback, were evaluated under field conditions in three experiments during the canola cropping seasons in Australia (April to November); two experiments: experiment-I (rainfed, 35 °03′36.9″S 147 °18′40.2″E), experiment-II (irrigated, 35 °02′27.0″S 147 °19′12.6″E) in 2013 at the WWAI, and one experiment: experiment-III (rainfed, 36.773949, 142.273688) at Green Lake, Victoria in 2015.
For the field experiments at Wagga Wagga, 169 DH lines along with the parental lines, and 12 commercial varieties of B. napus as checks [ATR-Marlin, ATR-Stingray, AV-Garnet, CB-Tanami, CB-Telfer, CB-Trigold, Crusher-TT, Hurricane-TT, Hyola50, Hyola76, Tawriffic-TT and one accession of Brassica juncea (Dune)] were grown as single rows (10 m long) in field blackleg screening nurseries, and spaced 80 cm apart in a randomised complete block design with three replications. The rows were trimmed to 8 m at growth stage 50 when plants reached the rosette stage. The nursery was planted on previous canola stubble that had been sprayed-out with glyphosate prior to seed set. The plants in the preceding stubble crop were a mixture of various genotypes with different known R genes in order to provide a diverse pathogen population for the subsequent season. The experiment-II trial was flood-irrigated (approximately 50 mm) once in September 2013 at the stem elongation stage to promote disease development. Trials were conducted as per standard agronomic practice, fertilized with 150 kg N, and sprayed with appropriate herbicides to control weeds when required.
For the experiment-III trial at Green Lake, 165 DH lines of RADH along with RP04, Ag-Outback and 22 B. napus commercial cultivars (44C79, 44Y84CL, 45Y86CL, ATR-Gem, ATR-Marlin, ATR-Stingray, CB-Sturt HT, CB-Telfer, Crusher TT, Hyola444TT, Hyola-50, Hyola 559TT, Hyola 575CL, JBOT800407, Nuseed Diamond, Pioneer 44Y26, Pioneer 44Y89, SturtTT, ThumperTT, Trigold, Victory V3002, and Westar) and two B. juncea cv. Dune and OasisCL as checks were sown in situ in the previous year’s canola stubble of cultivar ATR-GEM (Rlm1) on 28 May 2015. The trial was designed in AGROBASE using the ‘Grid Nursery with Checks’ model, which is a complete randomised block, with single row and two replicate design, and has a check cv. Trigold inserted every 10th row throughout the trial. The data collected from the disease nursery were: emergence counts at 6 weeks after sowing, adult plant survival counts and internal infection levels, both at physiological maturity. Up to 20 randomly selected plants from each row were cut from the crowns and scored for percent area discoloration of the stem due to blackleg fungus growth, as per scale described previously22.
Evaluation for resistance with ascospore shower test
171 DH lines from RADH and the parental lines, along with two controls; Skipton and Ag-Spectrum, were evaluated by Ag-Vic/Marcroft Grains Pathology for resistance to L. maculans pathotypes released from a mixed stubble using the ascopsore shower test23. Canola stubble was collected from commercial crops of AV-Garnet (MT Hope, South Australia), CB-Jardhee HT (Frances, South Australia), Monola76TT (Bool Lagoon, South Australia) and ATR-Cobbler (Wagga Wagga, NSW), grown in the 2011 cropping season. Details of the ‘ascospore shower’ test are given in Marcroft et al.22. This experiment was conducted in plastic pots (20 cm diameter) under glasshouse conditions in 2013 using a multiphase design comprising (i) an inoculation chamber phase and (ii) a glasshouse phase in order to account for non-genetic sources of variation in both phases. All entries after inoculation were laid in two chambers as shown in Supplementary Fig. 1. Each inoculation chamber accommodated a complete replicate of 200 entries (171 RADH lines, 2 entries of each parental lines; RP04 and Ag-Outback and 13 entries of Skipton and 12 entries of Ag-Spectrum) plus 50% of lines (100 lines) representing the third replicate. Each chamber had a complete array of 20 rows by 30 columns, ensuring that no two DH lines would be located next to each other across replicates. Four plants of each genotype were used as the experimental unit with each plant being scored. Spatial effects were modelled separately for both chambers. The design was generated using optimal design (od) software24 in R. At the physiological maturity stage (GS 83–85), the inoculated plants were cut at the crowns and cross sections were assessed visually for disease severity as described previously3.
Evaluation of RADH population for R genes
To determine whether R genes that confer resistance to L. maculans under controlled glasshouse conditions are the same with the ones contributing towards QR detected under field conditions, parental lines, RP04 and Ag-outback and their DH lines were characterized for resistance to L. maculans with 10 single-spore isolates (SSI) via cotyledon tests. Two isolates, IBCN17-D4 (AvrLm4, AvrLm5, AvrLm6, AvrLm7, AvrLm8, AvrLepR3, AvrLepR1, AvrLepR4) and IBCN76-D7 (AvrLm1, AvrLm3, AvrLm5, AvrLm6, AvrLm8, AvrLepR3, AvrLepR1, AvrLepR4)), that showed differential disease reactions on the parental lines (Supplementary Table 1a) were used for the genetic analysis of 177 DH lines of RADH in two independent experiments.
A randomized complete block design was generated using the spatial design search program DiGGer (Coombes 2002) assuming positive correlation between neighboring cells in rows and columns and allowing for random row and column effects within a column-pair of benches and further used for evaluating the DH population. A total of 171 DH lines, two parental lines and nine control cultivars; Westar (no R gene), AV-Garnet (Rlm1, 9), Q2 (Rlm3,9), Thunder TT (Rlm4), Caiman (Rlm7), Surpass400 (Rlm1 and LepR3), Hyola50 (Rlm1, LepR1), Mustang (Rlm6) and 44C79C (Rlm unknown) were evaluated in two replicates across two benches, each bench had a replicate of 23 trays, with each tray containing a row of 8 genotypes representing either DH and/or parental or control cultivars. The seedlings were grown in plastic trays (7 × 8 cells) in a controlled glasshouse at WWAI and maintained at 20 ± 2 °C for two weeks. Seven plants of each DH line (the experimental unit in a tray) were inoculated with the single spore isolate at a concentration of 106 spores mL−1 as described previously3. The inoculated plants were then placed in a humidity chamber for 48 h at 100% relative humidity in the dark at 18 °C to allow spore germination. Subsequently, inoculated plants were transferred to the glasshouse and maintained at 20 ± 3 °C for 3–4 weeks. Disease symptoms on cotyledons were scored 21 days post-inoculation using a scale described previously25: 0 (resistant response/no disease development) to 9 (highly susceptible/and profuse sporulation). Quantitative disease scores were used to identify significant trait-marker associations.
Phenotypic data analysis
Linear mixed model technology was used to fit a statistical model to phenotypic data generated by each experiment. The statistical model for each experiment included terms synonymous with a partitioning of the total variation into genetic and non-genetic sources of variation. Genetic sources of variation were further divided into variation due to DH lines and non-DH lines (which includes parental lines and commercial cultivars). Non-genetic sources of variation included terms associated with design factors associated with each experiment and in the case of field trials may also have included terms associated with the modelling of field spatial variability26. A logistic transformation was used for plant scores, internal infection percentage and survival percentage so that the statistical models for these traits better satisfied an assumption of normality and constant variance. The statistical model for each trait/experiment combination was used to form DH line Empirical Best Linear Unbiased Predictions (EBLUP’s) which are used in Fig. 1 to illustrate the range in resistance to L. maculans; and to estimate a broad sense heritability (h2)27 associated with DH lines. All statistical models were fitted using the statistical software package “asreml”28 within the R computing environment29.
Distribution of disease scores in 177 doubled haploid lines from RP04/Ag-Outback evaluated for resistance to L. maculans under glasshouse using two single spore isolates; IBCN76 (A) and IBCN17 (B), field (C–E), and shade house conditions (‘ascospore shower’ test) (F). Disease severity was assessed as internal canker formation on a 0 to 100 scale (Raman et al. 2012). Disease lesions on cotyledons were measured 21 days post-inoculation. (C) Arrows indicate the mean disease scores of RP04 and Ag-Outback at 21 days post-inoculation.
DArTseq genotyping
Genomic DNA was isolated from young leaves of 2–3 week-old seedlings of parental and DH lines from the RADH population using a standard phenol-chloroform method. The DNA samples were sent to Diversity Arrays Technology Pty. Ltd. (DArT P/L, University of Canberra, Bruce, Australia) for genotyping. Both parental lines and their DH lines were analysed using a DArTseq marker approach, as detailed in Raman et al.30. DArTseq markers were assigned to chromosomes based on their physical positions on the B. napus cv ‘Darmor-bzh’ genome assembly19.
Linkage map construction and QTL analysis
DArTseq markers having <80% call rate, and <5% MAF were discarded prior to linkage analysis. A set of 18,851 DArTseq-SNPs and silico DArTs that showed segregation among DH lines was selected for genetic linkage map construction as described in31. QTL analysis involved taking the statistical models for phenotypic data and partitioning DH line genetic variation into variation due to putative QTL and polygenic variation. Putative QTL for resistance to blackleg disease were identified using a genome scan31,32 as well as the whole genome average interval mapping approaches33. In this study, we considered trait-marker associations as ‘significant’ having LOD scores ≥3 and ‘suggestive’ with LOD score ≥2 for all environments (experiments). However, researchers can make their own decisions on the usefulness of putative QTL when developing their own marker-assisted selection strategies. Trait-marker association was graphically represented using the Map Chart software34.
Physical mapping and identification of putative candidate R genes
Sequences of markers significantly associated with resistance to blackleg were aligned to the reference B. napus cv. Darmor-bzh genome with the default BLAT settings19. Physical positions of R genes were downloaded from the public B. napus genome database (http://www.genoscope.cns.fr). Comparisons were made between the physical positions of B napus disease R genes and significant ‘hits’ using Microsoft Excel software. Candidate genes which were mapped within 200 kb of significant SNPs were checked for their annotations in TAIR (https://www.arabidopsis.org/) and B. napus genome databases. Biological functions of those genes in relation to R and QR loci were described.
Results
Inheritance of field resistance
Significant genetic variation for resistance scored on the basis of internal canker formation was observed in field experiments (Table 1). The internal canker scores varied from 0.1 to 99.21% in the RADH population depending on the phenotyping environment (Table 1). Disease scores showed a continuous distribution for genetic variation in resistance among the DH lines of RADH grown under field conditions, suggesting that quantitative genes control resistance to L. maculans (Fig. 1). Internal canker scores were higher in the experiment at Green Lake compared to the Wagga Wagga site. Across different field experiments, canker scores showed moderate (44%) to high repeatability (broad sense heritability, h2 = 79%) except in the Green Lake experiment where h2 was low (12%). High h2 (61%) for resistance to L. maculans was observed among the DH lines evaluated with the ascospore shower test, suggesting that resistance was genetically controlled, and that this test is also suitable for identification of loci for resistance to blackleg (Table 1). The Pearson’s correlation coefficient analysis showed poor to moderate genetic correlation (r = 0.17 to 0.5) between disease scores across sites (Fig. 2). These results confirmed that phenotypic environment and/or specificity of host resistance to the diverse L. maculans population contributed significantly to the differential disease expression.
Inheritance of resistance to single-spore isolates (SSI) of L. maculans
The parental lines of the RADH population were characterized for race-specific resistance using two SSI of L. maculans. Disease reactions showed that RP04 was resistant to the IBCN17 isolate, showing no visible critical disease symptoms (cotyledon score: 2.62) and Ag-Outback was susceptible (cotyledon lesion score: 5.26) (Supplementary Table 1b). However, Ag-Outback showed a resistance response to the IBCN76 isolate (2.66), while RP04 showed a susceptible response (5.65). Based on the differential response of parental lines of RADH population, Ag-Outback and RP04 are predicted to harbor Rlm3 and Rlm4 genes, respectively, for resistance to L. maculans. The distributions of cotyledon scores in the lines inoculated with IBCN17 and IBCN76 isolates showed a nearly bimodal distribution in a RADH population, suggesting that resistance to L. maculans could be due to race-specific R genes (Fig. 1A,B). The h2 estimates for disease scores were high, ranging from 85% to 97% for the IBCN76 and IBCN17 isolates, respectively (Table 1). The mean scores of DH lines ranged from 2.29 to 5.56 with IBCN17, compared to 2.60 to 5.80 with IBCN76. Significant negative correlation (r = −0.80) between disease scores of DH lines assessed with IBCN17 (AvrLm4, AvrLm5, AvrLm6, AvrLm7, AvrLm8, AvrLepR3, AvrLepR1, AvrLepR4) and IBCN76 (AvrLm1, AvrLm3, AvrLm5, AvrLm6, AvrLm8, AvrLepR3, AvrLepR1, AvrLepR4) was observed, suggesting that resistance in the population is contributed by different R genes (Rlm3 and Rlm4) conferred by both parents having opposite allelic effects.
Genetic mapping of RADH population
The RADH population was assayed with DArTseq-SNP and presence-absence (in silico DArTseq) marker variants. A total of 18,851 polymorphisms representing 6,442 SNPs and 12,409 in silico DArT markers were identified which showed segregation in the RADH population. Genotypic data of the RADH lines are presented in Supplementary Table 2. These polymorphic markers could be grouped into 2,019 unique bins, each one representing a single locus (Supplementary Table 3). Marker density on A subgenome was higher compared to C subgenome: 1051 bin markers were mapped on the subgenome A, covering 1,374.66 cM, and 968 bin markers were mapped on the subgenome C, covering 1,242.08 cM. The number of marker loci ranged from 149 on chromosome A08 to 1,942 on chromosome C04. On an average, bin marker density per cM ranged from 0.57 on chromosome A09 to 1.5 on chromosome C06, with a genome-wide coverage of 0.77 markers per cM. A framework linkage map based on 2, 019 bin markers (Supplementary Table 3) was further utilized for QTL analysis to identify trait-marker associations.
Genetic analysis of DH population for resistance to blackleg
A total of 42 SNP marker associations (22 significant associations with LOD ≥ 3 and 20 suggestive associations with LOD ≥ 2) were identified using genome scan approach for resistance to L. maculans evaluated under (i) field disease nursery conditions, (ii) controlled conditions with SSI, and (iii) diverse pathotypes derived from mixed stubble using the ascospore shower test (Supplementary Table 4). The number of QTL identified ranged from 4 to 12 depending on the screening method/phenotyping environment/QTL detection method (Tables 2 and 3), and accounted for 1.2% to 90.1% of the observed genetic variation. Whole genome average interval mapping (WGAIM) identified 26 genomic regions (QTL) for resistance to L. maculans on the 13 chromosomes of B. napus (Table 3).
Genetic mapping for field resistance
Three experiments were conducted to assess resistance to L. maculans; two at Wagga Wagga and one at Green Lake. Up to 11 significant marker associations were identified using both genome scan and WGAIM approaches on chromosomes A01, A03, A05, A06, A07, A10, C04, C06 and C08 in the RADH population (Tables 2 and 3). Interestingly, two QTL on chromosomes A03 and C04 were repeatedly detected across experiments, suggesting that environment; including prevalence of diverse races of L. maculans may have contributed toward inconsistent detection of QTLs. The percentage of genotypic variation accounted by individual SNP markers ranged from 3.4% (C01) to 33.3% (A10, Table 3). Among the significant SNPs, 3147873_8:C > T/4121514 marker interval explained the maximum genetic variation (33.3%) for resistance on chromosome A10.
Genetic mapping for resistance with ascospore shower test
Nine marker intervals showed significant associations for resistance to mixed L. maculans pathotypes originated from stubble sources, on chromosomes: A01, A03, A07, A09, A10, C2 C03, and C04 (Table 3). The genotypic variation explained by these QTL ranged from 2.8% to 18.1%. DArTseq SNP marker interval 4108895_33:A > G/3196349_56:A > C showed highly significant association (LOD = 9.36) on chromosome A01 and accounted for 12.4% of genetic variation in resistance to L. maculans, followed by marker interval 100027433_25:T > C/3098300_24:G > T with LOD score of 8.17 on chromosome A01, which accounted for 18.1% of genetic variation (Table 3). Collectively these QTL accounted for 48.76% of QR in RADH population. Both parents contributed favorable alleles for QR.
Mapping of race-specific resistance to L. maculans
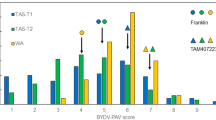
Six significant associations were detected for resistance to SSIs of L. maculans using genome scan approach; two SNP associations were detected with IBCN17 isolate on chromosomes A07 and A09, while four were detected with isolate IBCN76 on chromosomes A07, C02, C03 and C05 (Table 2). WGAIM detected two QTL regions for resistance to SSIs on chromosomes A07 and C09 (Table 3). The genotypic variation explained by individual SNP ranged from 2.8% to 90.1%. Highly significant marker interval 3076956-16:G > A/3096079_47:A > T (LOD = 90.0) on chromosome A07 accounted for 90.1% of the genetic variation for resistance to isolate IBCN17. An additive effect was also contributed by the maternal parent, RP04 (Tables 2 and 3). Several markers (20) showed complete segregation with DArTseq SNP marker 3076956-16:G > A on A07 (Supplementary Table 2), suggesting that these markers can be used for selection of resistance in different genetic backgrounds. The same marker interval; 3076956_16:G > A/3096079_47:A > T also showed significant association (LOD = 97.76) for resistance to isolate IBCN76, accounting for up to 80.8% of the genetic variation (Table 3, Fig. 3). However, an additive effect was also contributed by the paternal parent, Ag-Outback. This differential avirulent response of Ag-Outback to isolate IBCN76 confirmed the similar allelic effect revealed by the QTL mapping approaches. Given that SNP markers (3076956_16:G > A and 3096079_47:A > T on A07) accounted for a large proportion of genotypic variation for resistance to L. maculans (Table 3, Fig. 3), the disease scores from cotyledon tests were binned into two discrete categories; lines having average scores of ≤3 were classified as resistant and lines that have ≥4 were rated as susceptible. Linkage analysis revealed that a major locus for resistance to L. maculans; Rlm3 is located on chromosome A07 (Table 3). Linkage analysis between phenotypic binned data from cotyledon tests and markers present on a physical map of chromosome A07 of the reference genome showed that both the Rlm3 and Rlm4 genes are located on chromosome A07 (Fig. 3). The location of both R genes determined here is consistent with the previous mapping studies conducted on populations from Maxol/Westar, Columbus/Westar, Ag-Castle/Westar, and Ag-Castle/Topas3,5,10.
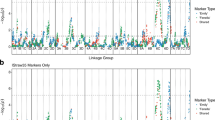
Partial genetic linkage maps of chromosomes A03, A07 and C04 of B. napus DH population from RP04/Ag-Outback, showing localization of Rlm3, Rlm4 genes (in rectangle) as well as QTL for resistance to L. maculans. Physical map positions of significant associations are shown on the reference sequence of B. napus v4.1.
Consistent QTL detected for resistance
Of the 19 significant QTLs identified with WGAIM, the allelic effects of the A03, A07 and C04 loci were detected repeatedly at least in two phenotypic environments (Table 3). The genetic effect of the A03 and C04 loci was repeatedly detected under field conditions at Wagga Wagga sites. Other consistent QTL on chromosome A07 for resistance to L. maculans evaluated by ascospore shower test was localized within the marker-intervals for Rlm3 and Rlm4. QTL for field resistance on chromosome A01 was also identified within 1 Mb of previously identified QTL for QR in B. napus populations5,35.
Identification of putative candidate resistance genes
To obtain the physical positions of putative R genes, 19 DArTseq SNP markers that showed significant association (LOD ≥ 3) with R and QR loci in the RADH population (Table 4), we searched for their identities against the reference sequence of Darmor using BLASTN. Only 60% of marker sequences were mapped on the reference sequence (Supplementary Table 1), suggesting that some of the genomic variation in B. napus varieties (e.g. RP04 and Ag-Outback) is not represented in the reference Darmor genome. Several markers were mapped on ChrAnn_ran contigs which have not been anchored on the reference B. napus.
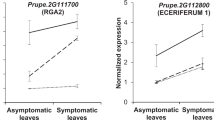
Physical mapping of markers linked with consistent resistance loci on the ‘Darmor’ genome assembly enabled identification of three candidate genes: BnaA03g26760D (A03), BnaA07g20240D (A07), and BnaC03g04240D (C03) (Table 4). These genes were searched for their identities against A. thaliana genome (TAIR 10 protein database). The BnaA03g26760D gene showed significant sequence identities (669 bits, E value = 0) with Transcription factor JUMONJI (AT4G00990.1), while the BnaA07g20240D (A07) gene showed sequence identities with Protein kinase superfamily protein of A. thaliana (AT1G79640.1; score bits: 1234, e value: 0.0). This gene has been previously mapped close to the Rlm4 locus for blackleg resistance36. BnaC03g04240D (C03) showed sequence similarities with AT5G10370.1 gene encoding helicase domain-containing protein (score: 2806, e value: 0.0). JUMONJI has been shown specifically regulated in the resistant SA1306 genotype suggesting that it act as major regulators of transcription defense responses against Erysiphe pisi in Medicago37. JUMONJI C has also been implicated in defense response against Xanthomomas oryzae pv. oryzae infection38.
Discussion
Virulent strains of L. maculans constantly threaten the sustainability of canola industry worldwide. Therefore, identification and deployment of both effective R and QR genes is needed for stable canola production. In previous studies, genetic resistance to L. maculans has been mapped in various Australian populations derived from Ag-Castle, AG-Spectrum, AV-Sapphire, Dunkeld, Maluka, Rainbow, Skipton, Shiralee and Surpass4003,4,5,6,8,9,10,39,40,41. Both parental lines of the RADH population, RP04 and Ag-Outback, have shown good levels of blackleg resistance under field conditions previously. In this study, we investigated the genetic basis of resistance and identified QTL for resistance to L. maculans in RADH population across six phenotypic environments. Among 11 significant SNP associations for field resistance, only two of them were detected repeatedly across experiments in both 2013 and 2015. This suggested that detection of QR (multi-genic) loci that are effective in conferring resistance to L. maculans across diverse environments is a challenging exercise. Highly heterogeneous populations of this pathogen42 may have contributed towards inconsistent detection of significant associations across environments (Tables 2 and 3). Secondly, experiments were conducted in different agro-climatic conditions which may have contributed to the different level of disease expression. For example in Lake Green, disease scores of DH lines had very low repeatability (12%), although canker scores were high (Table 1). In order to reduce the environmental variation and to increase selection efficiency for resistance, we utilized the ‘ascospore shower’ test to identify disease resistance loci under shade-house conditions. At least one QTL on chromosome A07 corresponding to the RP04 was detected with IBCN17 and ascospore shower test. This suggested that ascospore test is suitable for detecting both R and QR genes for resistance to L. maculans. Our results show that heritability increased from 12% (field experiment-site III) to 61% (under shade-house conditions, Table 1), suggesting that the ascospore shower can be reliably used for identification of QTL associated with resistance to blackleg pathotypes. In addition, a highly significant genetic association for resistance to L. maculans pathotypes was identified on chromosome A01 in RADH population. In previous studies, significant associations for resistance on A01 were reported in DH populations from Skipton/Ag-Spectrum, Topas/Ag-Castle and Topas/AV-Sapphire and diverse panels of B. napus3,5,10,17. Our results have also shown that race-specific major genes; Rlm3 and Rlm4 are ineffective at conferring resistance to L. maculans in RADH under Australian field conditions, as no QTL in the vicinity of Rlm3/Rlm4 genomic regions was identified on chromosome A07 (Table 4). It is probable that Rlm3- and Rlm4-attacking isolates were present in the phenotyping environments used, rendering both R genes ineffective. Our results are consistent with a previous study which shows that Rlm3 and Rlm4 are not very effective in providing resistance to L. maculans under Australian field conditions22. We identified 4 to 6 QTL associated with resistance using SSI (Table 3). While A07 loci represents to Rlm3 and Rlm4 genes, the role of other QTL on chromosomes A09, C2, C3, and C5 in conferring resistance to L. maculans need to be established. Since single spore isolates have multiple avirulence genes, it is possible that these loci may either (i) represent specific specificities, consistent with the gene-for-gene interaction (Flor 1971) in the B. napus-L. maculans pathosystem, or (ii) are genes triggered in response to pathogen attack (signaling and transduction). In B. napus, QR to L. maculans has been mapped onto chromosomes A09, A10, C02, C03 and C053,5,10.
In conclusion, we found that two race-specific genes Rlm3, Rlm4 and 19 significant QTLs control resistance to L. maculan in the DH population derived from the RP04/Ag-Outback cross. However, many stable QTLs could not be detected across environments as the QTL detection relies heavily on the phenotyping environment. QTL identified across multiple environments (years/sites) can be incorporated for enhancing effective QR in canola breeding germplasm. Although, the majority of QTL had small genetic effects, the genomic selection strategy could be used to combine a large number of environment-specific and environment non-specific (stable) QTL to develop varieties with durable resistance. Our study also showed that both Rlm3 and Rlm4 genes are ineffective in providing resistance under Australian field conditions. Therefore other strategies such as rotation of race-specific genes as well as deployment of stable quantitative resistance are required for sustainable blackleg management and canola production. Overall our results provide an insight into the R and QR loci for resistance to L. maculans in RP04/Ag-Outback.
References
Ansan-Melayah, D. et al. Genes for race-specific resistance against blackleg disease in Brassica napus L. Plant. Breed. 117, 373–378, https://doi.org/10.1111/j.1439-0523.1998.tb01956.x (1998).
Larkan, N. J. et al. The Brassica napus blackleg resistance gene LepR3 encodes a receptor-like protein triggered by the Leptosphaeria maculans effector AvrLM1. N. Phytologist 197, 595–605, https://doi.org/10.1111/nph.12043 (2013).
Raman, R. et al. Molecular mapping of qualitative and quantitative loci for resistance to Leptosphaeria maculans; causing blackleg disease in canola (Brassica napus L.). Theor. Appl. Genet. 125, 405–418, https://doi.org/10.1007/s00122-012-1842-6 (2012).
Raman, R. et al. Molecular mapping and validation of Rlm1 genes for resistance to Leptosphaeria maculans in canola (Brassica napus L). Crop. Pasture Sci. 63, 1007–1017, https://doi.org/10.1071/CP12255 (2012).
Raman, H. et al. Genome-wide association study identifies new loci for resistance to Leptosphaeria maculans in canola. Front. Plant. Sci. 7, 1513, https://doi.org/10.3389/fpls.2016.01513 (2016).
Raman, H. et al. Stable Quantitative Resistance Loci to Blackleg Disease in Canola (Brassica napus L.) Over Continents. Frontiers in Plant Science 9, https://doi.org/10.3389/fpls.2018.01622 (2018).
Huang, Y. J. et al. Identification of environmentally stable QTL for resistance against Leptosphaeria maculans in oilseed rape (Brassica napus). TAG. Theor. Appl. Genetics. Theoretische Und Angew. Genetik 129, 169–180, https://doi.org/10.1007/s00122-015-2620-z (2016).
Kaur, S. et al. Genetic map construction and QTL mapping of resistance to blackleg (Leptosphaeria maculans) disease in Australian canola (Brassica napus L.) cultivars. Theor. Appl. Genet. 120, 71–83 (2009).
Raman, H., Raman, R. & Larkan, N. Genetic Dissection of Blackleg Resistance Loci in Rapeseed (Brassica napus L.). Plant Breeding from Laboratories to Fields (Ed. Prof. Sven Bode Andersen). ISBN 978-953-51-1090-3, http://www.intechopen.com/articles/show/title/genetic-dissection-of-blackleg-resistance-loci-in-rapeseed-brassica-napus-l- <http://www.intechopen.com/books/export/citation/EndNote/plant-breeding-from-laboratories-to-fields/genetic-dissection-of-blackleg-resistance-loci-in-rapeseed-brassica-napus-l-> (2013).
Larkan, N. J. et al. Multi-environment QTL studies suggest a role for cysteine-rich protein kinase genes in quantitative resistance to blackleg disease in Brassica napus. BMC Plant. Biol. 16, 1–16, https://doi.org/10.1186/s12870-016-0877-2 (2016).
Raman, H. et al. SNP markers-based map construction and genome-wide linkage analysis in Brassica napus. Plant. Biotechnol. J. 12, 851–860, https://doi.org/10.1111/pbi.12186 (2014).
Li, H., Sivasithamparam, K. & Barbetti, M. J. Breakdown of a Brassica rapa subsp. sylvestris single dominant blackleg resistance gene in B. napus rapeseed by Leptosphaeria maculans field isolates in Australia. Plant Disease 87, 752 (710.1094/PDIS.2003.1087.1096.1752A), https://doi.org/10.1094/PDIS.2003.87.6.752A (2003).
Rouxel, T., Willner, E., Coudard, L. & Balesdent, M. H. Screening and identification of resistance to Leptosphaeria maculans (stem canker) in Brassica napus accessions. Euphytica 133, https://doi.org/10.1023/a:1025597622490 (2003).
Van de Wouw, A. P. et al. Breakdown of resistance to the fungal disease, blackleg, is averted in commercial canola (Brassica napus) crops in Australia. Field Crop. Res. 166, 144–151 (2014).
Fopa Fomeju, B., Falentin, C., Lassalle, G., Manzanares-Dauleux, M. J. & Delourme, R. Homoeologous duplicated regions are involved in quantitative resistance of Brassica napus to stem canker. BMC Genomics 15, 498, https://doi.org/10.1186/1471-2164-15-498 (2014).
Jestin, C. et al. Association mapping of quantitative resistance for Leptosphaeria maculans in oilseed rape (Brassica napus L.). Mol. Breed. 27, 271–287 (2011).
Rahman, M. et al. Association mapping in Brassica napus (L.) accessions identifies a major QTL for blackleg disease resistance on chromosome A01. Mol. Breed. 36, 1–15, https://doi.org/10.1007/s11032-016-0513-8 (2016).
Alamery, S. et al. Genome-wide identification and comparative analysis of NBS-LRR resistance genes in Brassica napus. Crop and Pasture. Sci. 69, 72–93, https://doi.org/10.1071/CP17214 (2018).
Chalhoub, B. et al. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345, https://doi.org/10.1126/science.1260782 (2014).
Cowling, W. A. Genetic diversity in Australian canola and implications for crop breeding for changing future environments. Field Crop. Res. 104, 103–111 (2007).
Cowley, R. B., Luckett, D. J., Moroni, J. S., Zeleke, K. & Diffey, S. Remote sensing of early vigour in canola germplasm: relationship to grain yield and potential to select for drought tolerance. Crop. Pasture Sci. 65(12), 1288–1299, https://doi.org/10.1071/CP14055 (2014).
Marcroft, S. J. et al. Identifying resistance genes to Leptosphaeria maculans in Australian Brassica napus cultivars based on reactions to isolates with known avirulence genotypes. Crop. Pasture Sci. 63, 338–350, https://doi.org/10.1071/CP11341 (2012).
Huang, Y. J. et al. Fitness cost associated with loss of the AvrLm4 avirulence function in Leptosphaeria maculans (Phoma stem canker of oilseed rape). Eur. J. Plant. Pathol. 114, 77–89 (2006).
Butler, D. G. od: An optimal design companion to ASReml. R Package version 0.71 (2011).
Koch, E., Song, K., Osborn, T. C. & Williams, P. H. Relationship between pathogenicity based on restriction fragment length polymorphism in Leptosphaeria maculans. Mol. Plant-Microbe Interact. 4, 341–349 (1991).
Gilmour, A. R., Cullis, B. R. & Verbyla, A. x16b & nas, P. Accounting for Natural and Extraneous Variation in the Analysis of Field Experiments. J. Agricultural, Biological, Environ. Stat. 2, 269–293, https://doi.org/10.2307/1400446 (1997).
Cullis, B. R., Smith, A. B. & Coombes, N. E. On the design of early generation variety trials with correlated data. J. Agricultural, Biological, Environ. Stat. 11, 381–393 (2006).
Butler, D. G., Cullis, B. R., Gilmour, A. R. & Gogel, B. J. ASReml-R reference manual. Release 3.0. Technical report, Queensland Department of Primary Industries, Australia, http://www.vsni.co.uk/downloads/asreml/release2/doc/asreml-R.pdf. (2009).
Team, R. D. C. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0, http://www.R-project.org/ (2014).
Raman, H. et al. Genome-Wide Delineation of Natural Variation for Pod Shatter Resistance in Brassica napus. PLoS ONE 9, e101673, https://doi.org/10.1371/journal.pone.0101673 (2014).
Raman, R. et al. Quantitative genetic analysis of yield in an Australian Brassica napus doubled haploid population. Crop & Pasture. Sci. 67, 298–307 (2016).
Nelson, M. N. et al. Quantitative Trait Loci for Thermal Time to Flowering and Photoperiod Responsiveness Discovered in Summer Annual-Type Brassica napus L. PLOS ONE 9, e102611, https://doi.org/10.1371/journal.pone.0102611 (2014).
Taylor, J. & Verbyla, A. R. Package wgaim: QTL Analysis in bi-parental populations using linear mixed models. Journal of Statistical Software 40(7), 1–18, http://www.jstatsoft.org/v40/i07/ (2011).
Voorrips, R. E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 93, 77–78, https://doi.org/10.1093/jhered/93.1.77 (2002).
Kumar, V. et al. Multi-year linkage and association mapping confirm the high number of genomic regions involved in oilseed rape quantitative resistance to blackleg. TAG. Theoretical and applied genetics. Theoretische und angewandte Genetik, https://doi.org/10.1007/s00122-018-3103-9 (2018).
Mayerhofer, R. et al. Complexities of chromosome landing in a highly duplicated genome: Toward map-based cloning of a gene controlling blackleg resistance in Brassica napus. Genet. 171, 1977–1988 (2005).
Curto, M., Krajinski, F., Schlereth, A. & Rubiales, D. Transcriptional profiling of Medicago truncatula during Erysiphe pisi infection. Front. Plant. Sci. 6, 517, https://doi.org/10.3389/fpls.2015.00517 (2015).
Hou, Y. et al. JMJ704 positively regulates rice defense response against Xanthomonas oryzae pv. oryzae infection via reducing H3K4me2/3 associated with negative disease resistance regulators. BMC Plant. Biol. 15, 1–13, https://doi.org/10.1186/s12870-015-0674-3 (2015).
Dusabenyagasani, M. & Fernando, W. G. D. Development of a SCAR marker to track canola resistance against blackleg caused by Leptosphaeria maculans pathogenicity group 3. Plant. Dis. 92, 903–908 (2008).
Mayerhofer, R., Bansal, V. K., Thiagarajah, M. R., Stringam, G. R. & Good, A. G. Molecular mapping of resistance to Leptosphaeria maculans in Australian cultivars of Brassica napus. Genome 40, 294–301 (1997).
Rimmer, S. R., Borhan, M. H., Zhu, B. & Somers, D. Mapping resistance genes in Brassica napus to Leptosphaeria maculans Proc. 10th Int. Rapeseed Congr. Canberra, Australia (1999).
Hayden, H. L., Cozijnsen, A. J. & Howlett, B. J. Microsatellite and minisatellite analysis of Leptosphaeria maculans in Australia reveals regional genetic differentiation. Phytopathology 97, 879–887, https://doi.org/10.1094/phyto-97-7-0879 (2007).
Acknowledgements
The authors thank Mr. David Roberts and several casual staff members of the NBGIP team for sowing research trials at the WWAI. This research was supported by the Grains Research and Development Corporation, NSWDPI, Ag Victoria and SAGI under projects DAN00117, DAV0085 and SAGI.
Author information
Authors and Affiliations
Contributions
H.R. and R.R. planned and carried out research work, S.D. and N.C. analysed the phenotypic data, H.R., R.R. and S.D. analysed trait-marker associations, D.L. and R.C. supported field trials in 2013, S.M. performed the ascospore shower test, D.B. and P.S. provided the seed of RP04/Ag-Outback DH population and evaluated for resistance at Green Lake. All authors commented and approved this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Raman, R., Diffey, S., Barbulescu, D.M. et al. Genetic and physical mapping of loci for resistance to blackleg disease in canola (Brassica napus L.). Sci Rep 10, 4416 (2020). https://doi.org/10.1038/s41598-020-61211-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-61211-y
This article is cited by
-
Pathogen lifestyle determines host genetic signature of quantitative disease resistance loci in oilseed rape (Brassica napus)
Theoretical and Applied Genetics (2024)
-
Epidemiology and management of blackleg of canola in response to changing farming practices in Australia
Australasian Plant Pathology (2021)
-
Status and advances in mining for blackleg (Leptosphaeria maculans) quantitative resistance (QR) in oilseed rape (Brassica napus)
Theoretical and Applied Genetics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.