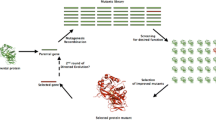
Abstract
Structural-genetic characterization of protease producing genes and enzymes from microbial sources are seldom appreciated despite having its substantial utilization in protein engineering or genetic manipulation for biotechnological applications. Aeromonas veronii CMF, a mesophilic bacterium isolated from the gut of Chrysomya megacephala, was found to exhibited significant level of protease activity. For the revelation of genetic potential in relation to protease production, whole genome of this organism was sequenced and analysed while structure–function of different protease enzyme was predicated using various in silico analysis. The 4.5 mb CMF genome was found to encompass various types of protease and mostly they are neutral in nature. Enzyme production was highest in an optimum pH and temperature of 6.0 (32.09 ± 1.015 U/ml) and 35ºC (41.65 ± 1.152 U/ml), respectively. Other culture parameters for optimum production of protease were determined to be inoculum size (1%), incubation period (72 h), shaking condition (125 rpm), carbon and nitrogen source [2% lactose (92.21 ± 3.16 U/ml) and 0.5% urea (163.62 ± 4.31 U/ml), respectively] and effect of surfactants [0.02 mg/ml Tween 80 (174.72 ± 4.48 U/ml)]. Furthermore, A. veronii CMF exhibited significant enzyme production like serine protease (15.22 ± 0.563 U/ml), aspartate protease (33.16 ± 0.762 U/ml) and collagenase (17.26 ± 0.626 U/ml). Genomic information and results of physio-biochemical assays indicate its cost-effective potential use in different enzyme-industry.






source and surfactants on protease production by CMF

Similar content being viewed by others
References
Abusham RA, Rahman RN, Salleh AB, Basri M (2009) Optimization of physical factors affecting the production of thermo-stable organic solvent-tolerant protease from a newly isolated halo tolerant Bacillus subtilis strain Rand. Microb Cell Fact 8:20
Adinarayana K, Jyothi B, Ellaiah P (2005) Production of alkaline protease with immobilized cells of Bacillus subtilis PE-11 in various matrices by entrapment technique. AAPS PharmSciTech 6:E391–E397
Akiyama Y, Kihara A, Tokuda H, Ito K (1996) FtsH (HflB) is an ATP-dependent protease selectively acting on SecY and some other membrane proteins. J Biol Chem 271:31196–31201
Amoozegar MA, Fatemi AZ, Karbalaei-Heidari HR, Razavi MR (2007) Production of an extracellular alkaline metalloprotease from a newly isolated, moderately halophile, Salinivibrio sp. strain AF-2004. Microbiol Res 162:369–377
Bajaj BK, Sharma P (2011) An alkali-thermotolerant extracellular protease from a newly isolated Streptomyces sp. DP2. N Biotechnol 28:725–732
Banerjee S, Maiti TK, Roy RN (2017) Protease production by thermo-alkaliphilic novel gut isolate Kitasatospora cheerisanensis GAP 12.4 from Gryllotalpa africana. Biocatal Biotransformation 35:168–176
Banerjee S, Maity TK, Roy RN (2020) Production, purification, characterization of cellulase from Acinetobacter junii GAC 16.2, a novel cellulolytic gut isolate of Gryllotalpa africana and its effects on cotton fiber and sawdust. Ann Microbiol 70:1–6
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Becnel JJ (1997) Complementary techniques: preparations of entomopathogens and diseased specimens for more detailed study using microscopy. In: Lacey L (ed) Manual of techniques in insect pathology. Academic Press, San Diego, pp 337–353
Benkert P, Biasini M, Schwede T (2011) Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 27:343–350
Berasategui A, Shukla S, Salem H, Kaltenpoth M (2016) Potential applications of insect symbionts in biotechnology. Appl Microbiol Biotechnol 100:1567–1577
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242
Bhattacharya A, Saini V, Gupta A (2012) Novel application of Mahua (Madhuca sp.) flowers for augmented protease production from Aeromonas sp. S1. Nat Prod Commun 7:1359–1362
Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T, Kiefer F, Cassarino TG, Bertoni M, Bordoli L, Schwede T (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42(W1):W252–W258
Bieniossek C, Schalch T, Bumann M, Meister M, Meier R, Baumann U (2006) The molecular architecture of the metalloprotease FtsH. Proc Natl Acad Sci USA 103:3066–3071
Bochtler M, Hartmann C, Song HK, Bourenkov GP, Bartunik HD, Huber R (2000) The structures of HslU and the ATP-dependent protease HslU–HslV. Nature 403:800–805
Borhani MS, Etemadifar Z, Emtiazi G, Jorjani E (2017) A statistical approach for production improvement of a neutral protease from a newly isolated strain of Aeromonas Hydrophila. Iran J Sci Technol Trans A Sci 42:1771–1778
Carnahan AM, Joseph SW (2005) Aeromonadaceae. In: Brenner DJ, Krieg JT, Garrity GM (eds) The Proteobacteria, Part B, Bergey’s manual of systematic bacteriology. Springer, New York
Chu WH (2006) Optimization of extracellular alkaline protease production from species of Bacillus. J Ind Microbiol Biotechnol 34:241–245
Cupp-Enyard C (2008) Sigma’s non-specific protease activity assay-444 casein as a substrate. J Vis Exp 19:e899
Diermayr P, Kroll S, Klostermeyer H (1987) Influence of EDTA and metal ions on a metalloproteinase from Pseudomonas fluorescensbiotype I. Biol Chem Hoppe Seyler 368:57–62
Divakar K, Priya JD, Gautam P (2010) Purification and characterization of thermostable organic solvent-stable protease from Aeromonas veronii PG01. J Mol Catal B Enzym 66:311–318
Duarte AS, Correia A, Esteves AC (2016) Bacterial collagenases–a review. Crit Rev Microbiol 42:106–126
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Euzéby JP (1997) List of bacterial names with standing in nomenclature: a folder available on the internet. Int J Syst Evol Microbiol 47:590–592
Evans EC, Abdullahi A (2012) Effect of surfactant inclusions on the yield and characteristics of protease from Bacillus subtilis. Proc Rom Acad Series B. 2:108–112
Gasteiger E, Hoogland C, Gattiker A, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The proteomics protocols handbook, Springer protocols handbooks. Humana Press, Totowa, pp 571–607
Ghareib M, Fawzi EM, Aldossary NA (2013) Thermostable alkaline protease from Thermomyces lanuginosus: optimization, purification and characterization. Ann Microbiol 64:859–867
Guruprasad K, Reddy BB, Pandit MW (1990) Correlation between stability of a protein and its dipeptide composition: a novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Eng Des Sel 4:155–161
Hayashi A, Aoyagi H, Yoshimura T, Tanaka H (2007) Development of novel method for screening microorganisms using symbiotic association between insect (Coptotermes formosanus Shiraki) and intestinal microorganisms. J Biosci Bioeng 103:358–367
Hedstrom L (2002) Serine protease mechanism and specificity. Chem Rev 102:4501–4524
Ibrahim AS, Al-Salamah AA (2009) Optimization of media and cultivation conditions for alkaline protease production by alkaliphilic Bacillus halodurans. Res J Microbiol 4:251–259
Ikai A (1980) Thermostability and aliphatic index of globular proteins. J Biochem 88:1895–1898
Jacob MB, Gerstein MJ (1960) Hand book of microbiology. D. Van Nostrand Co., Inc, New Jersey, p 61
Karbalaei-Heidari HR, Amoozegar MA, Hajighasemi M, Ziaee AA, Ventosa A (2008) Production, optimization and purification of a novel extracellular protease from the moderately halophilic bacterium Halobacillus karajensis. J Ind Microbiol Biotechnol 36:21–27
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10:845
Kumar S, Stecher G, Li M et al (2018) MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol 35:1547–1549
Kurtz S, Phillippy A, Delcher AL, Smoot M, Shumway M, Antonescu C, Salzberg SL (2004) Versatile and open software for comparing large genomes. Genome Biol 5:12
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132
Laishram S, Pennathur G (2015) Purification and characterization of a membrane-unbound highly thermostable metalloprotease from Aeromonas Caviae. Arab J Sci Eng 41:2107–2116
Leung KY, Stevenson RM (1988) Characteristics and distribution of extracellular proteases from Aeromonas hydrophila. Microbiology 134:151–160
Lim DV, Jackson RJ, Pull-VonGruenigen CM (1993) Purification and assay of bacterial collagenases. J Microbiol Methods 18:241–253
Llorens C, Futami R, Renau G, Moya A (2009) Bioinformatic flowchart and database to investigate the origins and diversity of Clan AA peptidases. Biol Direct 4:3
Lovell SC, Davis IW, Arendall WB et al (2003) Structure validation by Calpha geometry: phi, psi and Cbeta deviation. Proteins 50:437e450
Mahanta N, Gupta A, Khare SK (2008) Production of protease and lipase by solvent tolerant Pseudomonas aeruginosa PseA in solid-state fermentation using Jatropha curcas seed cake as substrate. Bioresour Technol 99:1729–1735
Mandal V, Sen SK, Mandal NC (2013) Production and partial characterization of an inducer-dependent novel antifungal compound(s) by Pediococcus acidilactici LAB 5. J Sci Food Agric 93:2445–2253
Mandal S, Rameez MJ, Chatterjee S, Sarkar J, Pyne P, Bhattacharya S, Shaw R, Ghosh W (2020) Molecular mechanism of sulfur chemolithotrophy in the betaproteobacterium Pusillimonas ginsengisoli SBSA. Microbiology 166:386–397
Mandal S, Debnath U, Sarkar J (2021) Structural-genetic characterization of novel butaryl co-a dehydrogenase and proposition of butanol biosynthesis pathway in Pusillimonas ginsengisoli SBSA. J Mol Evol 89:81–94
Matkawala F, Nighojkar S, Kumar A, Nighojkar A (2019) Enhanced production of alkaline protease by Neocosmospora sp. N1 using custard apple seed powder as inducer and its application for stain removal and dehairing. Biocatal Agric Biotechnol. 21:101310
Moldoveanu SC, David V, Moldoveanu SC, David V (2017) Properties of analytes and matrices determining HPLC Selection. Selection of the HPLC Method in Chemical Analysis, Elsevier, 189–230. https://doi.org/10.1016/B978-0-12-803684-6.00005-6
Moran NA (2006) Symbiosis. Curr Biol 16:866–871
Mothe T, Sultanpuram VR (2016) Production, purification and characterization of a thermotolerant alkaline serine protease from a novel species Bacillus caseinilyticus. 3 Biotech. 6:53
Oh YS, Shih L, Tzeng YM, Wang SL (2000) Protease produced by Pseudomonas aeruginosa K-187 and its application in the deproteinization of shrimp and crab shell wastes. Enzyme Microb Technol. https://doi.org/10.1016/S0141-0229(99)00172-6
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055
Parte AC (2014) LPSN—list of prokaryotic names with standing in nomenclature. Nucleic Acids Res 42:D613–D616
Patel R, Dodia M, Singh SP (2005) Extracellular alkaline protease from a newly isolated haloalkaliphilic Bacillus sp.: production and optimization. Process Biochem 40:3569–3575
Pushpam PL, Rajesh T, Gunasekaran P (2011) Identification and characterization of alkaline serine protease from goat skin surface metagenome. AMB Express 1:3
Rameez MJ, Pyne P, Mandal S, Chatterjee S, Alam M, Bhattacharya S, Mondal N, Sarkar J, Ghosh W (2019) Two pathways for thiosulfate oxidation in the alphaproteobacterial chemolithotroph Paracoccus thiocyanatus SST. Microbiol Res 230:126345
Rao MB, Tanksale AM, Ghatge MS, Deshpande VV (1998) Molecular and biotechnological aspects of microbial proteases. Microbiol Mol Biol Rev 62:597–635
Razzaq A, Shamsi S, Ali A, Ali Q, Sajjad M, Malik A, Ashraf M (2019) Microbial proteases applications. Front Bioeng Biotechnol. https://doi.org/10.3389/fbioe.2019.00110
Rohrwild M, Coux O, Huang HC, Moerschell RP, Yoo SJ, Seol JH, Chung CH, Goldberg AL (1996) HslV-HslU: a novel ATP-dependent protease complex in Escherichia coli related to the eukaryotic proteasome. Proc Natl Acad Sci USA 93:5808–5813
Saha S, Roy RN, Sen SK, Ray AK (2006) Characterization of cellulase-producing bacteria from the digestive tract of tilapia, Oreochromis mossambica (Peters) and grass carp, Ctenopharyngodon idella (Valenciennes). Aquac Res 37:380–388
Saini V, Bhattacharya A, Gupta A (2013a) Effectiveness of sal deoiled seed cake as an inducer for protease production from Aeromonas sp. S1 for its application in kitchen wastewater treatment. Appl Biochem Biotechnol 170:1896–1908. https://doi.org/10.1007/s12010-013-0323-y
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Singh J, Batra N, Sobti RC (2001) Serine alkaline protease from a newly isolated Bacillus sp. SSR1. Process Biochem 36:781–785
Souza PM, Werneck G, Aliakbarian B, Siqueira F, Ferreira Filho EX, Perego P, Converti A, Magalhães PO, Junior AP (2017) Production, purification and characterization of an aspartic protease from Aspergillus foetidus. Food Chem Toxicol 109:1103–1110
Stackebrandt E, Rainey FA (1995) Partial and complete 16S rDNA sequences, their use in generation of 16S rDNA phylogenetic trees and their implications in molecular ecological studies. In: Akkermans ADL, Van Elsas JD, De Bruijn FJ (eds) Molecular microbial ecology manual. Springer, Dordrecht, pp 259–275
Suganthi C, Mageswari A, Karthikeyan S, Anbalagan M, Sivakumar A, Gothandam KM (2013) Screening and optimization of protease production from a halotolerant Bacillus licheniformis isolated from saltern sediments. J Genet Eng Biotechnol 11:47–52
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA 101:11030–11035
Vega FE, Dowd PF (2005) The role of yeasts as insect endosymbionts. In: Blackwell M, Vega FE (eds) Insect-fungal associations: ecology and evolution. Oxford University Press, Oxford, pp 211–243
Waschkowitz T, Rockstroh S, Daniel R (2009) Isolation and characterization of metalloproteases with a novel domain structure by construction and screening of metagenomic libraries. Appl Environ Microbiol 75:2506–2516
Wen Z, Pengcheng D, Han Z et al (2014) Whole-genome sequence comparison as a method for improving bacterial species definition. J General Appl Microbiol 60(2):75–78
Yadav PK, Singh G, Gautam B et al (2013) Molecular modelling, dynamics studies and virtual screening of Fructose 1, 6 biphosphate aldolase-II in community acquired-methicillin resistant Staphylococcus aureus (CA-MRSA). Bioinformation 9:158
Zacaria J, Delamare AP, Costa SO, Echeverrigaray S (2010) Diversity of extracellular proteases among Aeromonas determined by zymogram analysis. J Appl Microbiol 1:212–219
Acknowledgements
We thank Dr. Wriddhiman Ghosh (Bose Institute, India) for his constructive and intellectual suggestions in manuscript preparation and for providing all kind of computational facilities. Financial support for conducting the microbiological studies was provided by Visva Bharati University via departmental revenue budget grants. R.B. and J.S. received fellowships from Council of Scientific and Industrial Research, GoI. S.B is thankful to Department of Biotechnology, Govt. of India for granting DBT Twinning Project [No. BT/PR25738/NER/95/1329/2017 dated December 24, 2018].
Funding
There was no external funding source for this work.
Author information
Authors and Affiliations
Contributions
NCM conceived the study, designed the experiments, interpreted the results and wrote the paper in conjunction with SM. Microbiological experiments and data analysis performed by RB, SM, SB, KS, JS and DB. Bioinformatics analysis performed by SM while SB and KS performed all biochemical experiments. All authors read and vetted the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
We declare that we have no conflicts of interest in publishing this work.
Human or animal participants
No humans or animals were used in this project.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bhattacherjee, R., Mandal, S., Banerjee, S. et al. Structural-genetic insight and optimization of protease production from a novel strain of Aeromonas veronii CMF, a gut isolate of Chrysomya megacephala. Arch Microbiol 203, 2961–2977 (2021). https://doi.org/10.1007/s00203-021-02282-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02282-x