Abstract
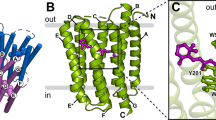
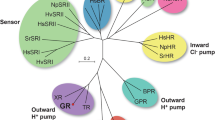
Bacteriorhodopsin is a transmembrane protein that uses light energy, absorbed by its chromophore retinal, to pump protons from the cytoplasm of bacteria such as Halobacterium salinarium into the extracellular space1,2. It is made up of seven α-helices, and in the bacterium forms natural, two-dimensional crystals called purple membranes. We have analysed these crystals by electron cryo-microscopy to obtain images of bacteriorhodopsin at 3.0 å resolution. The structure covers nearly all 248 amino acids, including loops outside the membrane, and reveals the distribution of charged residues on both sides of the membrane surface. In addition, analysis of the electron-potential map produced by this method allows the determination of the charge status of these residues. On the extracellular side, four glutamate residues surround the entrance to the proton channel, whereas on the cytoplasmic side, four aspartic acids occur in a plane at the boundary of the hydrophobic–hydrophilic interface. The negative charges produced by these aspartate residues is encircled by areas of positive charge that may facilitate accumulation and lateral movement of protons on this surface.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Oesterhelt, D. & Stoeckenius, W. Rhodopsin-like protein from the purple membrane of Halobacterium halobium. Nature New Biol. 233, 149–152 (1971).
Oesterhelt, D. & Stoeckenius, W. Functions of a new photoreceptor membrane. Proc. Natl Acad. Sci. USA 70, 2853–2857 (1973).
Khorana, H. G. Bacteriorhodopsin, a membrane protein that uses light to translocate protons. J. Biol. Chem. 263, 7439–7442 (1988).
Lanyi, J. K. Proton translocation mechanism and energetics in the light-driven pump bacteriorhodopsin. Biochim. Biophys. Acta 1183, 241–261 (1993).
Henderson, R. et al. Amodel for the structure of bacteriorhodopsin based on high resolution electron cryo-microscopy. J. Mol. Biol. 213, 899–929 (1990).
Grigorieff, N., Ceska, T. A., Downing, K. H., Baldwin, J. M. & Henderson, R. Electron-crystallographic refinement of the structure of bacteriorhodopsin. J. Mol. Biol. 259, 393–421 (1996).
Fujiyoshi, Y. et al. Development of a superfluid helium stage for high-resolution electron microscopy. Ultramicroscopy 38, 241–251 (1991).
Sakata, K., Tahara, Y., Morikawa, K., Fujiyoshi, Y. & Kimura, Y. Amethod for observing cross-sectional views of biomembranes. Ultramicroscopy 45, 253–261 (1992).
Gerwert, K., Hess, B., Soppa, J. & Oesterhelt, D. The role of 96 Asp in proton translocation by bacteriorhodopsin. Proc. Natl Acad. Sci. USA 86, 4943–4947 (1989).
Otto, H. et al. Aspartic acid-96 is the internal proton donor in the reprotonation of the Schiff base of bacteriorhodopsin. Proc. Natl Acad. Sci. USA 86, 9228–9232 (1989).
Braiman, M. S. et al. Vibrational spectroscopy of bacteriorhodopsin mutants: light-driven proton transport involves protonation changes of aspartic acid residues 85, 96, and 212. Biochemistry 27, 8516–8520 (1988).
Kimura, Y. & Ikegami, A. Local dielectric properties around polar region of lipid bilayer membranes. J. Membr. Biol. 85, 225–231 (1985).
Brown, L. S. et al. Glutamic acid 204 is the terminal proton release group at the extracellular surface of bacteriorhodopsin. J. Biol. Chem. 270, 27122–27126 (1995).
Kushwaha, S. C., Kates, M. & Stoeckenius, W. Comparison of purple membrane from Halobacterium cutirubrum and Halobacterium halobium. Biochim. Biophys. Acta 426, 703–710 (1976).
Subramaniam, S., Greenhalgh, D. A. & Khorana, H. G. Aspartic acid 85 in bacteriorhodopsin functions both as proton acceptor and negative counterion to the Schiff base. J. Biol. Chem. 267, 25730–25733 (1992).
Riesle, J., Oesterhelt, D., Dencher, N. A. & Heberle, J. D38 is an essential part of the proton translocation pathway in bacteriorhodopsin. Biochemistry 35, 6635–6643 (1996).
Mogi, T., Stern, L. J., Marti, T., Chao, B. H. & Khorana, H. G. Aspartic acid substitutions affect proton translocation by bacteriorhodopsin. Proc. Natl Acad. Sci. USA 85, 4148–4152 (1988).
Oesterhelt, D. & Stoeckenius, W. Isolation of the cell membrane of Halobacterium halobium and its fractionation into red and purple membrane. Methods Enzymol. 31, 667–678 (1974).
Seiff, F., Wallat, I., Ermann, P. & Heyn, M. Aneutron diffraction study on the location of the polyene chain of retinal in bacteriorhodopsin. Proc. Natl Acad. Sci. USA 82, 3227–3231 (1985).
Baldwin, J. & Henderson, R. Measurement and evaluation of electron diffraction patterns from two-dimensional crystals. Ultramicroscopy 14, 319–333 (1984).
Ceska, T. A. & Henderson, R. Analysis of high-resolution electron diffraction patterns from purple membrane labeled with heavy atoms J. Mol. Biol. 213, 539–560 (1990).
Collaborate Computational Project No. 4 Acta Crystallogr. D 50, 760–763 (1994).
International Union of Crystallography International table for Crystallography Volume C: Mathematical, Physical and Chemical Table (corrected edn) (ed. Wilson, A. J. C.) (Kluwer, Dordrecht, 1995).
Acknowledgements
We thank M. Ikehara for encouragement and support; W. Kühlbrandt and D. N. Wang for help with data processing; R. Henderson, S. Fuller, J. Lanyi, W. Stoeckenius, Y. Harada and J.Sasaki for helpful discussions; T. Miyata for preparing this manuscript; and Digital Equipment Corporation for help with computers.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kimura, Y., Vassylyev, D., Miyazawa, A. et al. Surface of bacteriorhodopsin revealed by high-resolution electron crystallography. Nature 389, 206–211 (1997). https://doi.org/10.1038/38323
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/38323
This article is cited by
-
Regulation of membrane protein structure and function by their lipid nano-environment
Nature Reviews Molecular Cell Biology (2023)
-
X-ray structure analysis of bacteriorhodopsin at 1.3 Å resolution
Scientific Reports (2018)
-
Light-Patterned Current Generation in a Droplet Bilayer Array
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.