Abstract
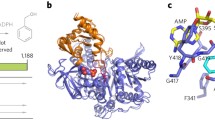
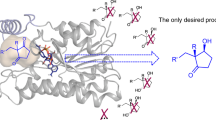
Automated structural analysis of Sporobolomyces salmonicolor carbonyl reductase (SSCR) indicates that the two largest potential receptor sites are in the vicinity of the nicotinamide reductant. The largest receptor site is a scalene triangle with sides of ∼8 Å by 9 Å by 13 Å, which is narrow in width; one corner is surrounded by hydrophilic residues that can favorably bond with the ketone oxygen. Docking aryl alkyl ketones shows a distinct preference for binding to the largest receptor site, and for conformations that place the carbonyl oxygen of the substrate in the hydrophilic corner of the largest receptor site. Favorable docking conformations for aryl alkyl ketones fall into two low-energy ensembles. These conformational ensembles are distinguished by the positions of the substituents, presenting either the Si-or Re-face of the ketone to the nicotinamide reductant. For the ketones investigated here, there is a correspondence between the major enantiomer of the alcohol obtained from the reduction of the ketone and the conformer found to have the most stable interaction energy with the receptor site in all cases. The receptor site modeling, docking simulations, molecular dynamics, and enzyme-substrate geometry optimizations lead to a model for understanding the enantioselectivity of this NADPH-dependent carbonyl reductase.

Receptor site model for NADPH-dependent carbonyl reductase





Similar content being viewed by others
References
Faber K (2004) Biotransformation in organic chemistry. Springer, Berlin Heidelberg New York
Ema T, Yagasaki H, Okita N, Takeda M, Sakai T (2006) Tetrahedron 62:6143–6149
Edegger K, Stampfer W, Seisser B, Faber K, Mayer SF, Oehrlein R, Hafner A, Kroutil W (2006) Eur J Org Chem 1904–1909
Poessl TM, Kosjek B, Ellmer U, Gruber CC, Edegger K, Faber K, Hildebrandt P, Bornscheuer UT, Kroutil W (2005) Adv Synth Catal 347:1827–1834
Ema T, Yagasaki H, Okita N, Nishikawa K, Korenaga T, Sakai T (2005) Tetrahedron: Asymmetry 16:1075–1078
Soni P, Kaur G, Chakraborti AK, Banerjee UC (2005) Tetrahedron: Asymmetry 16:2425–2428
Kaluzna IA, Matsuda T, Sewell AK, Stewart JD (2004) J Am Chem Soc 126:12827–12832
van Deursen R, Stampfer W, Edegger K, Faber K, Kroutil W (2004) J Mol Catal B: Enzymatic 31:159–163
Zhu D, Mukherjee C, Rozzell JD, Kambourakis S, Hua L (2006) Tetrahedron 62:901–905
Zhu D, Rios BE, Rozzell JD, Hua L (2005) Tetrahedron: Asymmetry 16:1541–1546
Zhu D, Mukherjee C, Hua L (2005) Tetrahedron: Asymmetry 16:3275–3278
Kita K, Fukura T, Nakase K-I, Okamoto K, Yanase H, Kataoka M, Shimizu S (1999) Appl Environ Microbiol 65:5207–5211
Zhu D, Yang Y, Buynak JD, Hua L (2006) Org Biomol Chem 4:2690–2695
Zhu D, Hua L (2006) J Org Chem: 71:9484–9486
Kamitori S, Iguchi A, Ohtaki A, Yamada M, Kita K (2005) J Mol Biol 352:551–558
Chemical Computing Group, Inc., http://www.chemcomp.com
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM Jr, Ferguson DM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) J Am Chem Soc 117:5179–5197
Gasteiger J, Marsili M (1980) Tetrahedron 36:3219–3222
Liang J, Edelsbrunner H, Fu P, Sudhakar PV, Subramaniam S (1998) Proteins 33:1–17
Liang J, Edelsbrunner H, Fu P, Sudhakar PV, Subramaniam S (1998) Proteins 33:18–29
Baxter CA, Murray CW, Clark DE, Westhead DR, Eldridge MD (1998) Proteins 33:367–382
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) Nucleic Acids Res 28:235–242
Acknowledgements
A.D. and T.R.C. acknowledge the U.S. Department of Education for their support of CASCaM. These authors gratefully acknowledge the Chemical Computing Group for generously providing the Molecular Operating Environment (MOE) program. D. Z. and L. H. thank Southern Methodist University for generous financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cundari, T.R., Dinescu, A., Zhu, D. et al. A molecular modeling study on the enantioselectivity of aryl alkyl ketone reductions by a NADPH-dependent carbonyl reductase. J Mol Model 13, 685–690 (2007). https://doi.org/10.1007/s00894-007-0168-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-007-0168-9