Abstract
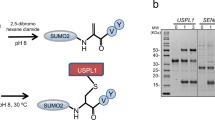
Small ubiquitin-like modifier (SUMO)-specific protease SENP1 processes SUMO-1, SUMO-2 and SUMO-3 to mature forms and deconjugates them from modified proteins. To establish the proteolytic mechanism, we determined structures of catalytically inactive SENP1 bound to SUMO-1–modified RanGAP1 and to unprocessed SUMO-1. In each case, the scissile peptide bond is kinked at a right angle to the C-terminal tail of SUMO-1 and has the cis configuration of the amide nitrogens. SENP1 preferentially processes SUMO-1 over SUMO-2, but binding thermodynamics of full-length SUMO-1 and SUMO-2 to SENP1 and Km values for processing are very similar. However, kcat values differ by 50-fold. Thus, discrimination between unprocessed SUMO-1 and SUMO-2 by SENP1 is based on a catalytic step rather than substrate binding and is likely to reflect differences in the ability of SENP1 to correctly orientate the scissile bonds in SUMO-1 and SUMO-2.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Meluh, P.B. & Koshland, D. Evidence that the MIF2 gene of Saccharomyces cerevisiae encodes a centromere protein with homology to the mammalian centromere protein CENP-C. Mol. Biol. Cell 6, 793–807 (1995).
Rosas-Acosta, G., Russell, W.K., Deyrieux, A., Russell, D.H. & Wilson, V.G. A universal strategy for proteomic studies of SUMO and other ubiquitin-like modifiers. Mol. Cell. Proteomics 4, 56–72 (2005).
Vertegaal, A.C. et al. A proteomic study of SUMO-2 target proteins. J. Biol. Chem. 279, 33791–33798 (2004).
Holmstrom, S., Van Antwerp, M.E. & Iniguez-Lluhi, J.A. Direct and distinguishable inhibitory roles for SUMO isoforms in the control of transcriptional synergy. Proc. Natl. Acad. Sci. USA 100, 15758–15763 (2003).
Azuma, Y., Arnaoutov, A. & Dasso, M. SUMO-2/3 regulates topoisomerase II in mitosis. J. Cell Biol. 163, 477–487 (2003).
Bernier-Villamor, V., Sampson, D.A., Matunis, M.J. & Lima, C.D. Structural basis for E2-mediated SUMO conjugation revealed by a complex between ubiquitin-conjugating enzyme Ubc9 and RanGAP1. Cell 108, 345–356 (2002).
Lin, D. et al. Identification of a substrate recognition site on ubc9. J. Biol. Chem. 277, 21740–21748 (2002).
Sampson, D.A., Wang, M. & Matunis, M.J. The small ubiquitin-like modifier-1 (SUMO-1) consensus sequence mediates Ubc9 binding and is essential for SUMO-1 modification. J. Biol. Chem. 276, 21664–21669 (2001).
Melchior, F., Schergaut, M. & Pichler, A. SUMO: ligases, isopeptidases and nuclear pores. Trends Biochem. Sci. 28, 612–618 (2003).
Tatham, M.H. et al. Polymeric chains of sumo-2 and sumo-3 are conjugated to protein substrates by sae1/sae2 and ubc9. J. Biol. Chem. 276, 35368–35374 (2001).
Li, S.J. & Hochstrasser, M. A new protease required for cell-cycle progression in yeast. Nature 398, 246–251 (1999).
Mossessova, E. & Lima, C.D. Ulp1-SUMO crystal structure and genetic analysis reveal conserved interactions and a regulatory element essential for cell growth in yeast. Mol. Cell 5, 865–876 (2000).
Yeh, E.T., Gong, L. & Kamitani, T. Ubiquitin-like proteins: new wines in new bottles. Gene 248, 1–14 (2000).
Gan-Erdene, T. et al. Identification and characterization of DEN1, a deneddylase of the ULP family. J. Biol. Chem. 278, 28892–28900 (2003).
Mendoza, H.M. et al. NEDP1, a highly conserved cysteine protease that deNEDDylates Cullins. J. Biol. Chem. 278, 25637–25643 (2003).
Wu, K. et al. DEN1 is a dual function protease capable of processing the C-terminus of Nedd8 deconjugating hyper-neddylated CUL1. J. Biol. Chem. 278, 28882–28891 (2003).
Di Bacco, A. et al. The SUMO-specific protease SENP5 is required for cell division. Mol. Cell. Biol. 26, 4489–4498 (2006).
Gong, L. & Yeh, E.T. Characterization of a family of nucleolar sumo-specific proteases with preference for sumo-2 or sumo-3. J. Biol. Chem. 281, 15869–15877 (2006).
Yamaguchi, T. et al. Mutation of SENP1/SuPr-2 reveals an essential role for desumoylation in mouse development. Mol. Cell. Biol. 25, 5171–5182 (2005).
Gong, L., Millas, S., Maul, G.G. & Yeh, E.T. Differential regulation of sentrinized proteins by a novel sentrin-specific protease. J. Biol. Chem. 275, 3355–3359 (2000).
Shen, L.N., Dong, C., Liu, H., Naismith, J.H. & Hay, R.T. The structure of SENP1 SUMO-2 co-complex suggests a structural basis for discrimination between SUMO paralogues during processing. Biochem. J. 397, 279–288 (2006).
Xu, Z. & Au, S.W. Mapping residues of SUMO precursors essential in differential maturation by SUMO-specific protease, SENP1. Biochem. J. 386, 325–330 (2005).
Reverter, D. & Lima, C.D. Insights into E3 ligase activity revealed by a SUMO-RanGAP1-Ubc9-Nup358 complex. Nature 435, 687–692 (2005).
Reverter, D. et al. Structure of a complex between Nedd8 and the Ulp/Senp protease family member Den1. J. Mol. Biol. 345, 141–151 (2005).
Shen, L.N. et al. Structural basis of NEDD8 ubiquitin discrimination by the deNEDDylating enzyme NEDP1. EMBO J. 24, 1341–1351 (2005).
Xu, Z. et al. Crystal structure of the SENP1 mutant C603S-SUMO complex reveals the hydrolytic mechanism of SUMO specific protease. Biochem. J. 398, 345–352 (2006).
Baba, D. et al. Crystal structure of thymine DNA glycosylase conjugated to SUMO-1. Nature 435, 979–982 (2005).
Pichler, A. et al. SUMO modification of the ubiquitin-conjugating enzyme E2–25K. Nat. Struct. Mol. Biol. 12, 264–269 (2005).
Reverter, D. & Lima, C.D. A basis for SUMO protease specificity provided by analysis of human Senp2 and a Senp2-SUMO complex. Structure 12, 1519–1531 (2004).
Stryer, L. Fluorescence energy transfer as a spectroscopic ruler. Annu. Rev. Biochem. 47, 819–846 (1978).
Cheng, Y. & Prusoff, W.H. Relationship between the inhibition constant (K1) and the concentration of inhibitor which causes 50 per cent inhibition (I50) of an enzymatic reaction. Biochem. Pharmacol. 22, 3099–3108 (1973).
Johnston, S.C., Riddle, S.M., Cohen, R.E. & Hill, C.P. Structural basis for the specificity of ubiquitin C-terminal hydrolases. EMBO J. 18, 3877–3887 (1999).
Hu, M. et al. Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell 111, 1041–1054 (2002).
Macauley, M.S. et al. Structural and dynamic independence of isopeptide-linked RanGAP1 and SUMO-1. J. Biol. Chem. 279, 49131–49137 (2004).
Radisky, E.S. & Koshland, D.E.J. A clogged gutter mechanism for protease inhibitors. Proc. Natl. Acad. Sci. USA 99, 10316–10321 (2002).
Liu, B., Schofield, C.J. & Wilmouth, R.C. Structural analyses on intermediates in serine protease catalysis. J. Biol. Chem. 281, 24024–24035 (2006).
Klabunde, T., Sharma, S., Telenti, A., Jacobs, W.R., Jr. & Sacchettini, J.C. Crystal structure of GyrA intein from Mycobacterium xenopi reveals structural basis of protein splicing. Nat. Struct. Biol. 5, 31–36 (1998).
Brauer, A.B., Domingo, G.J., Cooke, R.M., Matthews, S.J. & Leatherbarrow, R.J. A conserved cis peptide bond is necessary for the activity of Bowman-Birk inhibitor protein. Biochemistry 41, 10608–10615 (2002).
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20, 87–90 (2002).
Tatham, M.H. et al. Role of an N-terminal site of Ubc9 in SUMO-1, -2, and -3 binding and conjugation. Biochemistry 42, 9959–9969 (2003).
Leslie, A.G.W. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 ESF-EAMCB Newsl. Prot. Crystallogr. 26, 1–110 (1992).
Collaborative Computational Project, No. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
McCoy, A.J., Grosse-Kunstleve, R.W., Storoni, L.C. & Read, R.J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D Biol. Crystallogr. 61, 458–464 (2005).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Bossis, G. et al. A fluorescence resonance energy transfer-based assay to study SUMO modification in solution. Methods Enzymol. 398, 20–32 (2005).
Acknowledgements
pHis-TEV-30a was a kind gift from H. Liu, University of St. Andrews. This work was supported by CRUK, AICR and the Wellcome Trust. Structural analysis was carried out in the Scottish Structural Proteomics Facility, which is funded by the Biotechnology Biological Science Research Council and The Scottish Funding Council. J.H.N. is a Biotechnology Biological Science Research Council career development fellow.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Final 2Fo – Fc density at scissile bonds. (PDF 282 kb)
Rights and permissions
About this article
Cite this article
Shen, L., Tatham, M., Dong, C. et al. SUMO protease SENP1 induces isomerization of the scissile peptide bond. Nat Struct Mol Biol 13, 1069–1077 (2006). https://doi.org/10.1038/nsmb1172
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1172
This article is cited by
-
Develop quantitative FRET (qFRET) technology as a high-throughput universal assay platform for basic quantitative biomedical and translational research and development
Med-X (2023)
-
SUMO promotes longevity and maintains mitochondrial homeostasis during ageing in Caenorhabditis elegans
Scientific Reports (2020)
-
The Epstein-Barr Virus Oncoprotein, LMP1, Regulates the Function of SENP2, a SUMO-protease
Scientific Reports (2019)
-
Viral manipulation of the cellular sumoylation machinery
Cell Communication and Signaling (2017)
-
A new trend to determine biochemical parameters by quantitative FRET assays
Acta Pharmacologica Sinica (2015)