Abstract
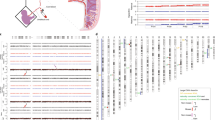
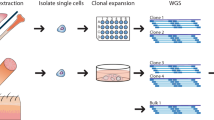
We demonstrate hre that somatic variation of CGG repeat length is based on a mosaic of cells with different but stable FMR–1 alleles and does not reflect permanent mitotic instability. The length of a particular allele in an individual cell was maintained in progeny cells establishing a clone. The mutation patterns of multiple repeats in the DNA of fetal tissues were identical and did not significantly change during proliferation in vitro. It is proposed that genotype mosaicism and expansion to full mutation are generated post–conceptionally by the same molecular mechanism in a particular window of early development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Vincent, A. et al. Abnormal pattern detected in fragile-X patients by pulsed-field gel electrophoresis. Nature 349, 624–626 (1991).
Oberlé, I. et al. Instability of a 550-base pair DNA segment and abnormal methylation in fragile X syndrome. Science 252, 1097–1102 (1991).
Yu, S. et al. Fragile X genotype characterized by an unstable region of DNA. Science 252, 1179–1181 (1991).
Bell, M.V. et al. Physical mapping across the fragile X: Hypermethylation and clinical expression of the fragile X syndrome. Cell 64, 861–866 (1991).
Verkerk, A.J.M.H. et al. Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 65, 905–914 (1991).
Kremer, E.J. et al. Mapping of DNA instability at the fragile X to a trinucletide repeat sequence p(CCG)n. Science 252, 1711–1714 (1991).
Fu, Y.-H. et al. Variation of the CGG repeat at the fragile X site results in genetic instability: Resolution of the Sherman paradox. Cell 67, 1047–1058 (1991).
Rousseau, F. et al. Direct diagnosis by DNA analysis of the fragile X syndrome of mental retardation. New Engl. J. Med. 325, 1673–1681 (1991).
Wöhrle, D. et al. Genotype mosaicism in fragile X fetal tissues. Hum. Genet. 89, 114–116 (1992).
Devys, D. et al. Analysts of full fragile X mutations in fetal tissues and monozygotic twins indicate that abnormal methylation and somatic heterogeneity are established early in development. Am. J. med. Genet. 43, 208–216 (1992).
Richards, R.I. & Sutherland, G.R. Fragile X syndrome: the molecular picture comes into focus. Trends Genet. 8, 249–254 (1992).
Smith, C.L., Klco, S.R. & Cantor, C.R. in Genome analysis, a practical approach (ed Davies, K. E.) 41–72 (IRL, 1988).
Wenger, S.L., Hennessy, J.C. & Steele, W. Increased sister chromatid exchange frequency at Xq27 site in affected fragile X males. Am. J. med. Genet. 26, 909 (1987).
Tommerup, N. Induction of the fragile X on BrdU-substituted chromosomes with direct visualization of sister chromatid exchanges on banded chromosomes. Hum. Genet. 81, 377–381 (1989).
Fonatsch, C. Fragile X chromosome in fibroblasts: Longitudinal study on technical variation. Clin. Genet. 23, 229 (1983).
Pieretti, M. et al. Absence of expression of the FMR-1 gene in fragile X syndrome. Cell 66, 817–822 (1991).
Rousseau, F. et al. Efficient and reliable direct diagnosis of the fragile X mental retardation syndrome. New Engl. J. Med. 325, 1673–1681 (1991).
Hori, T. et al. Heritable unstable DNA sequences and hypermethylation associated with fragile X syndrome in Japanese families. Clin. Genet. 43, 34–38 (1993).
Willems, P.J. et al. Segregation of the fragile X mutation from an affected male to his normal daughter. Hum. molec. Genet. 7, 511–515 (1992).
Wöhrle, D. & Steinbach, P. Fragile X expression and X inactivation. II. The fragile site at Xq27.3 has a basic function in the pathogenesis of fragile X-linked mental retardation. Hum. Genet. 87, 421–24 (1991).
Heitz, D., Devys, D., Imbert, G., Kretz, C. & Mandel, J.-L. Inheritance of the fragile X syndrome: Size of fragile X premutation is a major determinant of the transition to full mutation. J. med. Genet. 29, 794–801 (1992).
Crane, J.P. & Cheung, S.W. An embryonic model to explain cytogenetic inconsistencies observed in chorionic villus versus fetal tissue. Prenat. Diagn. 8, 119–129 (1988).
Takagi, N., Sugawara, O. & Sasaki, M. Regional and temporal changes in the pattern of X chromosome replication during early postimplantation development of the female mouse. Chromosoma 85, 275–286 (1982).
Nakahori, Y. et al. Molecular heterogeneity of the fragile X syndrome. Nucl. Acids Res. 19, 4355–4359 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wöhrle, D., Hennig, I., Vogel, W. et al. Mitotic stability of fragile X mutations in differentiated cells indicates early post–conceptional trinucleotide repeat expansion. Nat Genet 4, 140–142 (1993). https://doi.org/10.1038/ng0693-140
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0693-140
This article is cited by
-
CGG-repeat dynamics and FMR1 gene silencing in fragile X syndrome stem cells and stem cell-derived neurons
Molecular Autism (2016)
-
EMQN best practice guidelines for the molecular genetic testing and reporting of fragile X syndrome and other fragile X-associated disorders
European Journal of Human Genetics (2015)
-
Clinical utility gene card for: fragile X mental retardation syndrome, fragile X-associated tremor/ataxia syndrome and fragile X-associated primary ovarian insufficiency
European Journal of Human Genetics (2011)
-
Features of trinucleotide repeat instability in vivo
Cell Research (2008)
-
Post-zygotic de novo trinucleotide repeat expansion at spinocerebellar ataxia type 7 locus: evidence from an Indian family
Journal of Human Genetics (2005)