Abstract
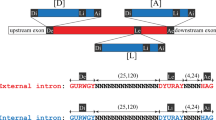
The nucleotides immediately surrounding intron/exon junctions of genes transcribed by RNA polymerase B can be derived from ‘consensus’ sequences for donor and acceptor splice sites by only a few base changes1–3. Studies in vivo have underlined the importance of these junction nucleotides for splicing4–7. In higher eukaryotes, no evidence has been found for specific internal intron sequences involved in splicing8–10. However, the recent discovery that, in vitro, introns are excised in a lariat form where the 5′ end of the intron is joined via a 2′-5′-phosphodiester linkage to an A residue (branchpoint acceptor) close to the 3′ end of the intron, suggests that internal intron sequences may nonetheless be important for splicing11–13. Indeed, in yeast nuclear genes, the internal sequence 5′-TACTAAC-3′ (or close homologue) is essential for splicing in vivo14,15. A proposed consensus sequence for branchpoints in mammalian introns is 5′-CT(A/G)A(C/T)-3′ (refs 11, 16, 17). This sequence resembles the essential yeast internal sequence. Are branchpoints involved in the splicing of introns of higher eukaryotes in vivo? We show here that a branchpoint sequence from a human globin gene (5′-CTGACTCTCTCTG-3′)11 greatly enhances the efficiency of splicing of a ‘synthetic’ intron in HeLa cells. A mutated branchpoint sequence, 5′-CTCCTCTCTCTG-3′, in which the branchpoint acceptor nucleotide A has been deleted and the neighbouring purine G mutated to a C, does not exhibit this enhancing capability. We conclude that branchpoints have an important function in the splicing process in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Breathnach, R., Benoist, C., O'Hare, K., Gannon, F. & Chambon, P. Proc. natn. Acad. Sci. U.S.A. 75, 4852–4857 (1978).
Breathnach, R. & Chambon, P. A. Rev. Biochem. 50, 349–383 (1981).
Mount, S. Nucleic Acids Res. 10, 465–473 (1982).
Wieringa, B., Meyer, F., Reiser, J. & Weissmann, C. Nature 301, 38–43 (1983).
Treisman, R., Orkin, S. H. & Maniatis, T. Nature 302, 591–596 (1983).
Mount, S. & Steitz, J. Nature 303, 380–381 (1983).
Rautmann, G., Matthes, H. W. D., Gait, M. J. & Breathnach, R. EMBO J. 3, 2021–2028 (1984).
Volkaert, G., Feunteun, J., Crawford, L. V., Berg, P. & Fiers, W. J. J. Virol 30, 677–682 (1979).
Wieringa, B., Hofer, E. & Weissmann, C. Cell 37, 915–925 (1984).
Dierks, P. et al. ICN UCLA Symp. molec. cell. Biol. 33, 347–366 (1981).
Ruskin, B., Krainer, A. R., Maniatis, T. & Green, M. R. Cell 38, 317–331 (1984).
Padgett, R. A., Konarski, M. M., Grabowski, P. J., Hardy, S. F. & Sharp, P. A. Science 225, 898–903 (1984).
Weissmann, C. Nature 311, 103–104 (1984).
Pikielny, C. W., Teem, J. L. & Rosbash, M. Cell 34, 395–403 (1984).
Langford, C. J. & Gallwitz, D. Cell 33, 519–527 (1983).
Keller, E. B. & Noon, W. A. Proc. natn. Acad. Sci. U.S.A. 81, 7417–7420 (1984).
Zeitlin, S. & Efstratiadis, A. Cell 39, 589–602 (1984).
Matthes, H. W. D. et al. EMBO J. 3, 801–805 (1984).
Tooze, J. (ed.) DNA Tumor Viruses 2nd edn (Cold Spring Harbor Laboratory, New York, 1982).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Rautmann, G., Breathnach, R. A role for branchpoints in splicing in vivo. Nature 315, 430–432 (1985). https://doi.org/10.1038/315430a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/315430a0
This article is cited by
-
Increased complexity of Tmem16a/Anoctamin 1 transcript alternative splicing
BMC Molecular Biology (2011)
-
The role of small nuclear ribonucleoprotein particles in pre-mRNA splicing
Nature (1987)
-
The expression of a nopaline synthase ? human growth hormone chimaeric gene in transformed tobacco and sunflower callus tissue
Plant Molecular Biology (1986)
-
Cleavage of 5′ splice site and lariat formation are independent of 3′ splice site in yeast mRNA splicing
Nature (1985)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.