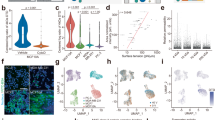
Abstract
We review a current methodology which analyzes quantitative scattering data from Scanning Transmission Electron Microscopy to establish footprints—positions of DNA-binding proteins on cloned genes—and mass distributions for the bound protein. Recognition by computer of the DNA path and the extent of the bound protein leads to an analysis rate which is high for image-based data. The methods have been applied in the study of eukaryotic transcription initiation and T antigen interaction with the SV40 control region.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Matsui, T., Segall, J., Weil, P.A. and Roeder, R.G. 1980. Multiple factors required for accurate initiation of transcription by purified RNA polymerase II. J. Biol. Chem. 255: 11,992–11,996.
Samuels, M., Fire, A. and Sharp, P.A. 1982. Separation and characterization of factors mediating accurate transcription by RNA polymerase II. J. Biol. Chem. 257: 14,419–14,427.
Dignam, J.D., Martin, P.L., Shastry, B.S. and Roeder, R.G. 1983. Eukaryotic gene transcription with purified components. Meth. Enzymol. 101: 583–598.
Davison, B.L., Egly, J.-M., Mulvihill, E.R. and Chambon, P. 1983. Formation of stable preinitiation complexes between eukaryotic class B transcription factors and promoter sequences. Nature 301: 680–686.
Parker, C.S. and Topol, J. 1984. A Drosophila RNA polymerase II transcription factor contains a promoter-region-specific DNA-binding activity. Cell 36: 357–369.
Wu, C. 1984. Two protein-binding sites in chromatin implicated in the activation of heat-shock genes. Nature 309: 229–234.
Gidoni, D., Dynan, W.S. and Tjian, R. 1984. Multiple specific contacts between a mammalian transcription factor and its cognate promoters. Nature 312: 409–413.
Fire, A., Samuels, M. and Sharp, P.A. 1984. Interactions between RNA polymerase II, factors, and template leading to accurate transcription. J. Biol. Chem. 259: 2509–2516.
Bogenhagen, D.F., Wormington, W.M. and Brown, D.D. 1982. Stable transcription complexes of Xenopus 5S RNA genes: a means to maintain the differentiated state. Cell 28: 413–421.
Lassar, A.B., Martin, P.L. and Roeder, R.G. 1983. Transcription of class III genes: formation of preinitiation complexes. Science 222: 740–748.
Brown, D.D. 1984. The role of stable complexes that repress and activate eucaryotic genes. Cell 37: 359–365.
Bieker, J.J., Martin, P.L. and Roeder, R.G. 1985. Formation of a rate-limiting intermediate in 5S RNA gene transcription. Cell 40: 119–127.
Hough, P.V.C., Mastrangelo, I.A., Wall, J.S., Hainfeld, J.F., Simon, M.N. and Manley, J.L. 1982. DNA-protein complexes spread on N2-discharged carbon film and characterized by molecular weight and its projected distribution. J. Mol. Biol. 160: 375–386.
Hough, P.V.C., Simon, M.N. and Mastrangelo, I.A. 1984. Analysis of eukaryotic control proteins at their recognition sequences by scanning transmission electron microscopy, p. 279–307. In: Genetic Engineering, Vol. 6. Hollaender A. and Setlow, J. K. (eds.), Plenum Publ. Corp., N.Y.
Mastrangelo, I.A., Hough, P.V.C., Wilson, V.G., Wall, J.S., Hainfeld, J.F. and Tegtmeyer, P. 1985. Monomers through trimers of large T antigen in region I and monomers through tetramers in region II bind to SV40 origin DNA as stable structures in solution. Proc. Natl. Acad. Sci. USA in press.
Wall, J.S. 1979. Biological scanning transmission electron microscopy, p. 333–342. In: Introduction to Analytical Electron Microscopy. Hren, J.J., Goldstein, J. I., and Joy, D. C. (eds.), Plenum Publ. Co., N.Y.
Mosesson, M.W., Hainfeld, J., Wall, J. and Haschemeyer, R.H. 1981. Identification and mass analysis of human fibrinogen molecules and their domains by scanning transmission electron microscopy. J. Mol. Biol. 153: 695–718.
Langmore, J.P., Wall, J. and Isaacson, M.S. 1973. The collection of scattered electrons in dark field electron microscopy. 1. Elastic scattering. Optik 38: 335–350.
Buchman, A.R., Burnett, L. and Berg, P. 1980. The SV40 nucleotide sequence, p. 799–818. In: DNA Tumor Viruses. Tooze, J. (ed.), Cold Spring Harbor Laboratory, N.Y.
Henderson, R. 1975. Structure of the purple membrane from Halobacterium halobrum: analysis of the x-ray diffraction pattern. J. Mol. Biol. 93: 123–138.
Wall, J.S., Hainfeld, J.F., Bartlett, P.A. and Singer, S.J. 1982. Observation of an undecagold cluster compound in the scanning transmission electron microscope. Ultramicros. 8: 397–402.
Hough, P.V.C. 1962. General purpose visual input for a computer. Ann. N.Y. Acad. Sci. 99: 323–334.
Weil, P.A., Luse, D.S., Segall, J. and Roeder, R.G. 1979. Selective and accurate initiation of transcription at the Ad2 major late promoter in a soluble system dependent on purified RNA polymerase II and DNA. Cell 18: 469–484.
Manley, J.L., Fire, A., Cano, A., Sharp, P.A. and Gefter, M.L. 1980. DNA-dependent transcription of adenovirus genes in a soluble whole-cell extract. Proc. Natl. Acad. Sci. USA 77: 3855–3859.
Dignam, J.D., Lebovitz, R.M. and Roeder, R.G. 1984. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11: 1475–1489.
Slattery, E., Dignam, J.D., Matsui, T. and Roeder, R.G. 1983. Purification and analysis of a factor which suppresses nick-induced transcription by RNA polymerase II and its identity with poly(ADP-ribose) polymerase. J. Biol. Chem. 258: 5955–5959.
Sawadogo, M. and Roeder, R.G. 1984. Energy requirement for specific transcription initiation by the human polymerase II system. J. Biol. Chem. 259: 5321–5326.
Tegtmeyer, P. 1975. Altered patterns of protein synthesis in infection by SV40 mutants. Cold Spring Harbor Symp. Quant. Biol. 39: 9–15.
Myers, R.M. and Tjian, R. 1980. Construction and analysis of simian virus origins defective in tumor antigen binding and DNA replication. Proc. Natl. Acad. Sci. USA 77: 6491–6495.
Shortle, D.R., Margolskee, R.F. and Nathans, D. 1979. Mutational analysis of the simian virus 40 replicon: pseudorevertants of mutants with a defective replication origin. Proc. Natl. Acad. Sci. USA 76: 6128–6131.
Wilson, V.G., Tevethia, M.J., Lewton, B.A. and Tegtmeyer, P. 1982. DNA-binding properties of simian virus 40 temperature-sensitive A proteins. J. Virol. 44: 458–468.
Hansen, U., Tenen, D.G., Livingston, D.M. and Sharp, P. 1981. T antigen repression of SV40 early transcription from two promoters. Cell 27: 603–612.
Rio, D.C. and Tjian, R. 1983. SV40 T antigen binding site mutations that affect autoregulation. Cell 32: 1227–1240.
Tjian, R. 1978. Protein-DNA interactions at the origin of simian virus 40 DNA replication. Cold Spring Harbor Symp. Quant. Biol. 43: 655–662.
DeLucia, A.L., Lewton, B.A., Tjian, R. and Tegtmeyer, P. 1983. Topography of simian virus A protein-DNA complexes: arrangement of pentanucleotide interaction sites at the origin of replication. J. Virol. 46: 143–150.
Tegtmeyer, P., Lewton, B.A., DeLucia, A.L., Wilson, V.G. and Ryder, K. 1983.Topography of simian virus 40 A protein-DNA complexes: arrangement of protein bound to the origin of replication. J. Virol. 46: 151–161.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hough, P., Mastrangelo, I., Wall, J. et al. STEM Footprints and Bound Mass Distributions for DNA Control Proteins. Nat Biotechnol 3, 549–553 (1985). https://doi.org/10.1038/nbt0685-549
Issue Date:
DOI: https://doi.org/10.1038/nbt0685-549