Abstract
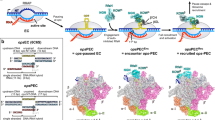
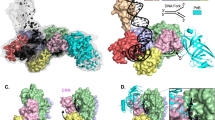
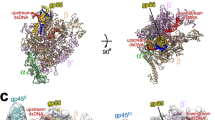
Plasmids are natural vectors for gene transfer. In Gram-negative bacteria, plasmid DNA replication is triggered when monomers of an initiator protein (Rep) bind to direct repeats at the origin sequence. Rep dimers, which are inactive as initiators, bind to an inverse repeat operator, repressing transcription of the rep gene. Rep proteins are composed of N-terminal dimerization and C-terminal DNA-binding domains. Activation of Rep is coupled to dimer dissociation, converting the dimerization domain into a second origin-binding module. Although the structure of the monomeric F plasmid initiator (mRepE) has been determined, the molecular nature of Rep activation remains unknown. Here we report the crystal structure of the dimeric N-terminal domain of the pPS10 plasmid initiator (dRepA). dRepA has a winged-helix fold, as does its homologous domain in mRepE. However, dimerization transforms an interdomain loop and β-strand (monomeric RepE) into an α-helix (dimeric RepA). dRepA resemble the C terminus of eukaryotic and archaeal Cdc6, giving clues to the phylogeny of DNA replication initiators.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
del Solar, G., Giraldo, R., Ruíz-Echevarría, M.J., Espinosa, M. & Díaz, R. Replication and control of circular bacterial plasmids. Microbiol. Mol. Biol. Rev. 62, 434–464 (1998).
Giraldo, R. Common domains in the initiators of DNA replication in Bacteria, Archaea and Eukarya: combined structural, functional and phylogenetic perspectives. FEMS Microbiol. Rev. 26, 533–554 (2003).
Manen, D., Upegui, G.L. & Caro, L. Monomers and dimers of the RepA protein in plasmid pSC101 replication: domains in RepA. Proc. Natl. Acad. Sci. USA 89, 8923–8927 (1992).
Ishiai, M., Wada, C., Kawasaki, Y. & Yura, T. Replication initiator protein RepE of mini-F plasmid: functional differentiation between monomers (initiator) and dimers (autogenous repressor). Proc. Natl. Acad. Sci. USA 91, 3839–3843 (1994).
García de Viedma, D., Giraldo, R., Ruíz-Echevarría, M.J., Lurz, R. & Díaz-Orejas, R. Transcription of repA, the gene of the initiation protein of the Pseudomonas plasmid pPS10, is autoregulated by sequential interactions of the RepA protein at a symmetrical operator. J. Mol. Biol. 247, 211–223 (1995).
Wickner, S., Skowyra, D., Hoskins, J. & McKenney, K. DnaJ, DnaK, and GrpE heat shock proteins are required in oriP1 DNA replication solely at the RepA monomerization step. Proc. Natl. Acad. Sci. USA 89, 10345–10349 (1992).
Sozhamannan, S. & Chattoraj, D.K. Heat shock proteins DnaJ, DnaK and GrpE stimulate P1 plasmid replication by promoting initiator binding to the origin. J. Bacteriol. 175, 3546–3555 (1993).
DasGupta, S., Mukhopadhyay, G., Papp, P.P., Lewis, M.S. & Chattoraj, D.K. Activation of DNA binding by the monomeric form of the P1 replication initiator RepA by heat shock proteins DnaJ and DnaK. J. Mol. Biol. 232, 23–34 (1993).
Wickner, S. et al. A molecular chaperone, ClpA, functions like DnaK and DnaJ. Proc. Natl. Acad. Sci. USA 91, 12218–12222 (1994).
Dibbens, J.A., Muraiso, K. & Chattoraj, D.K. Chaperone-mediated reduction of RepA dimerization is associated with RepA conformational change. Mol. Microbiol. 26, 185–195 (1997).
Konieczny, I. & Helinski, D.R. The replication initiator protein of the broad-host-range plasmid RK2 is activated by the ClpX chaperone. Proc. Natl. Acad. Sci. USA 94, 14378–14382 (1997).
Díaz-López, T. et al. Structural changes in RepA, a plasmid replication initiator, upon binding to origin DNA. J. Biol. Chem. 278, 18606–18616 (2003).
García de Viedma, D., Giraldo, R., Rivas, G., Fernández-Tresguerres, E. & Díaz-Orejas, R. A leucine-zipper motif determines different functions in a DNA replication protein. EMBO J. 15, 925–934 (1996).
Giraldo, R., Andreu, J.M. & Díaz-Orejas, R. Protein domains and conformational changes in the activation of RepA, a DNA replication initiator. EMBO J. 17, 4511–4526 (1998).
Nieto, C., Giraldo, R., Fernández-Tresguerres, E. & Díaz, R. Genetic and functional analysis of the basic replicon of pPS10, a plasmid specific of Pseudomonas isolated from Pseudomonas syringae pv. savastanoi. J. Mol. Biol. 223, 415–426 (1992).
Komori, H. et al. Crystal structure of a prokaryotic replication initiator protein bound to DNA at 2.6 Å resolution. EMBO J. 18, 4597–4607 (1999).
Gajiwala, K.S. & Burley, S.K. Winged helix proteins. Curr. Op. Struct. Biol. 10, 110–116 (2000).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 233, 123–138 (1993).
Giraldo, R., Nieto, C., Fernández-Tresguerres, M.E. & Díaz, R. Bacterial zipper. Nature 342, 866 (1989).
García de Viedma, D., Serrano-López, A. & Díaz-Orejas, R. Specific binding of the replication protein of plasmid pPS10 to direct and inverted repeats is mediated by an HTH motif. Nucleic Acids Res. 23, 5048–5054 (1995).
Liu, J. et al. Structure and function of Cdc6/Cdc18: implications for origin recognition and checkpoint control. Mol. Cell 6, 637–648 (2000).
Giraldo, R. & Díaz-Orejas, R. Similarities between the DNA replication initiators of Gram-negative bacteria plasmids (RepA) and eukaryotes (Orc4p)/archaea (Cdc6p). Proc. Natl. Acad. Sci. USA 98, 4938–4943 (2001).
Maestro, B., Sanz, J.M., Díaz-Orejas, R. & Fernández-Tresguerres, E. Modulation of pPS10 host range by plasmid-encoded RepA initiator protein. J. Bacteriol. 185, 1367–1375 (2003).
Matsunaga, F. et al. The central region of RepE initiator protein of mini-F plasmid plays a crucial role in dimerization required for negative replication control. J. Mol. Biol. 274, 27–38 (1997).
Xia, G., Manen, D., Yu, Y. & Caro, L. In vivo and in vitro studies of a copy number mutation of the RepA replication protein of plasmid pSC101. J. Bacteriol. 175, 4165–4175 (1993).
Miron, A., Patel, I. & Bastia, D. Multiple pathways of copy control of γ replicon of R6K: mechanisms both dependent on and independent of cooperativity of interaction of π protein with DNA affect the copy number. Proc. Natl. Acad. Sci. USA 91, 6438–6442 (1994).
Chattoraj, D.K. Control of plasmid DNA replication by iterons: no longer paradoxical. Mol. Microbiol. 37, 467–476 (2000).
Blasina, A., Kittell, B.L., Toukdarian, A.E. & Helinski, D.R. Copy-up mutants of the plasmid RK2 replication initiation protein are defective in coupling RK2 replication origins. Proc. Natl. Acad. Sci. USA 93, 3559–3564 (1996).
Urh, M. et al. Assemblies of replication initiation protein on symmetric and asymmetric DNA sequences depend on multiple protein oligomerization surfaces. J. Mol. Biol. 283, 619–631 (1998).
Uga, H., Matsunaga, F. & Wada, C. Regulation of DNA replication by iterons: an interaction between the ori2 and incC regions mediated by RepE-bound iterons inhibits DNA replication of mini-F plasmid in Escherichia coli. EMBO J. 18, 3856–3867 (1999).
Park, K., Han, E., Paulsson, J. & Chattoraj, D.K. Origin pairing ('handcuffing') as a mode of negative control of P1 plasmid copy number. EMBO J. 20, 7323–7332 (2001).
Spolar, R.S. & Record, T.M. Jr. Coupling of local folding to site-specific binding of proteins to DNA. Science 263, 777–784 (1994).
Jen-Jacobson, L., Engler, L.E. & Jacobson, L.A. Structural and thermodynamic strategies for site-specific DNA binding proteins. Structure 8, 1015–1023 (2000).
Leslie, A.G.W. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 ESF-EAMCB Newslett. Protein Crystallogr. 26 (1992).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
de La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
Laskowski, R.A, MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structure. J. Appl. Crystallogr. 26, 283–291 (1993).
Kraulis, P.J. MOLSCRIPT—a program to produce both detailed and schematic plots of protein structures. Appl. Crystallogr. 24, 946–950 (1996).
Merritt, E.A. & Bacon, D.J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank the staff at beamline X31 in EMBL-DESY for help with data collection. We are indebted to D. Rhodes for the critical reading of the manuscript and, together with V. Ramakrishnan, for encouragement. This work has been financed with grants of Spanish MCyT, CAM and a FIS network for research on infectious pathology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Giraldo, R., Fernández-Tornero, C., Evans, P. et al. A conformational switch between transcriptional repression and replication initiation in the RepA dimerization domain. Nat Struct Mol Biol 10, 565–571 (2003). https://doi.org/10.1038/nsb937
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb937
This article is cited by
-
Transcriptional expression of secondary resistance genes ccdB and repA2 is enhanced in presence of cephalosporin and carbapenem in Escherichia coli
BMC Microbiology (2021)
-
RepA-WH1, the agent of an amyloid proteinopathy in bacteria, builds oligomeric pores through lipid vesicles
Scientific Reports (2016)
-
Functional amyloids as inhibitors of plasmid DNA replication
Scientific Reports (2016)
-
Pre-amyloid oligomers of the proteotoxic RepA-WH1 prionoid assemble at the bacterial nucleoid
Scientific Reports (2015)
-
Plasmid pP62BP1 isolated from an Arctic Psychrobacter sp. strain carries two highly homologous type II restriction-modification systems and a putative organic sulfate metabolism operon
Extremophiles (2012)