Summary
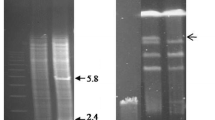
The structural gene, PHO13, for the specific p-nitrophenyl phosphatase of Saccharomyces cerevisiae was cloned and its nucleotide sequence determined. The deduced PHO13 protein consists of 312 amino acids and its molecular weight is 34635. The disruption of the PHO13 gene produced no effect on cell growth, sporulation, or viability of ascospores. The PHO13 locus was mapped at 1.9 centimorgans from the HO locus on the left arm of chromosome IV. By chromosome fragmentation, the PHO13 locus was found to be located about 72 kb from the left-hand telomere of chromosome IV and distal to the HO locus.
Similar content being viewed by others
References
Ammerer G, Hunter CP, Rothman JH, Saari GC, Valls LA, Stevens TH (1986) PEP4 gene of Saccharomyces cerevisiae encodes proteinase A, a vacuolar enzyme required for processing of vacuolar precursors. Mol Cell Biol 6:2490–2499
Attias J, Bonnet JL (1972) A specific alkaline p-nitrophenyphosphatase activity from baker's yeast. Biochim Biophys Acta 268:422–430
Attias J, Durand H (1973) Further characterization of a specific p-nitrophenylphosphatase from baker's yeast. Biochim Biophys Acta 321:561–568
Attias J, Bonnet JL, Sauvagnargues JC (1970) Séparation et étude partielle de deux phosphatases alcalines de levure. Biochim Biophys Acta 212:315–321
Bradshaw RA, Cancedda F, Ericsson LH, Neumann PA, Piccoli SP, Schlesinger MJ, Shriefer K, Walsh KA (1981) Amino acid sequence of Escherichia coli alkaline phosphatase. Proc Natl Acad Sci USA 78:3473–3477
Carle GF, Olson MV (1985) An electrophoretic karyotype for yeast. Proc Natl Acad Sci USA 82:3756–3760
Clark DW, Tkacz JS, Lampen JO (1982) Asparagine-linked carbohydrate does not determine the cellular location of yeast vacuolar nonspecific alkaline phosphatase. J Bacteriol 152:865–873
Clarke L, Carbon J (1978) Functional expression of cloned yeast DNA in Escherichia coli: specific complementation of argininosuccinate lyase (argH) mutations. J Mol Biol 120:517–532
Feinberg AP, Vogelstein B (1983) A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 132:6–13
Fujita M, Nakao T, Tashima Y, Mizuno N, Nagano K, Nakao M (1966) Potassium-ion stimulated p-nitrophenylphosphatase activity occurring in a highly specific adenosine triphosphatase preparation from rabbit brain. Biochim Biophys Acta 117:42–53
Gorman JA, Hu ASL (1969) The separation and partial characterization of l-histidinol phosphatase and an alkaline phosphatase of Saccharomyces cerevisiae. J Biol Chem 244:1645–1650
Harashima S, Takagi A, Oshima Y (1984) Transformation of protoplasted yeast cells is directly associated with cell fusion. Mol Cell Biol 4:771–778
Hieter P, Mann C, Snyder M, Davis RW (1985) Mitotic stability of yeast chromosomes: A colony color assay that measures nondisjunction and chromosome loss. Cell 40:381–392
Kaneko Y, Toh-e A, Oshima Y (1982) Identification of the genetic locus for the structural gene and a new regulatory gene for the synthesis of repressible alkaline phosphatase in Saccharomyces cerevisiae. Mol Cell Biol 2:127–137
Kaneko Y, Tamai Y, Toh-e A, Oshima Y (1985) Transcriptional and post-transcriptional control of PHO8 expression by PHO regulatory genes in Saccharomyces cerevisiae. Mol Cell Biol 5:248–252
Kaneko Y, Hayashi N, Toh-e A, Banno I, Oshima Y (1987) Structural characteristics of the PHO8 gene encoding repressible alkaline phosphatase in Saccharomyces cerevisiae. Gene 58:137–148
Knuuttila MLE, Mäkinen KK (1972) Purification and characterization of a phosphatase specifically hydrolyzing p-nitrophenyl phosphate from an oral strain of Streptococcus mutans. Arch Biochem Biophys 152:685–701
Langford CJ, Gallwitz D (1983) Evidence for an intron-contained sequence required for the splicing of yeast RNA polymerase lI transcripts. Cell 33:519–527
Lehle L, Bause E (1984) Primary structural requirements for Nand O-glycosylation of yeast mannoproteins. Biochim Biophys Acta 799:246–251
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: A laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Matsumoto K, Uno I, Kato K, Ishikawa T (1985) Isolation and characterization of a phosphoprotein phosphatase-deficient mutant in yeast. Yeast 1:25–38
Maxam AM, Gilbert W (1980) Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol 65:499–560
Mitchell JK, Fonzi WA, Wilkerson J, Opheim DJ (1981) A particulate form of alkaline phosphatase in the yeast, Saccharomyces cerevisiae. Biochim Biophys Acta 657:482–494
Mortimer RK, Schild D (1985) Genetic map of Saccharomyces cerevisiae, edition 9. Microbiol Rev 49:181–212
Oshima Y (1982) Regulatory circuits for gene expression, the metabolism of galactose and phosphate. In: Strathern JN, Jones EW, Broach JR (eds) The molecular biology of the yeast Saccharomyces: metabolism and gene expression. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, pp 159–180
Pallen CJ, Wang JH (1983) Calmodulin-stimulated dephosphorylation of p-nitrophenyl phosphate and free phospbotyrosine by calcineurin. J Biol Chem 258:8550–8553
Parent SA, Fenimore CM, Bostian KA (1985) Vector systems for the expression, analysis and cloning of DNA sequences in S. cerevisiae. Yeast 1:83–138
Rao GJS, Del Monte M, Nadler HL (1971) A K+-activated, ethacrynic acid-sensitive p-nitrophenylphosphatase from normal human white cells. Biochim Biophys Acta 229:454–457
Sherman F, Fink GR, Hicks JB (1986) Methods in yeast genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Shortle D, Haber JE, Botstein D (1982) Lethal disruption of the yeast actin gene by integrative DNA transformation. Science 217:371–373
Southern EM (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98:503–517
Stewart AA, Ingebritsen TS, Manalan A, Klee CB, Cohen P (1982) Discovery of a Ca2+- and calmodulin-dependent protein phosphatase, probable identity with calcineurin (CaM-BP80). FEBS Lett 137:80–84
Swarup G, Cohen S, Garbers DL (1981) Selective dephosphorylation of proteins containing phosphotyrosine by alkaline phosphatases. J Biol Chem 256:8197–8201
Toh-e A, Nakamura H, Oshima Y (1976) A gene controlling the synthesis of non-specific alkaline phosphatase in Saccharomyces cerevisiae. Biochim Biophys Acta 428:182–192
Vollrath D, Davis RW, Connelly C, Hieter P (1988) Physical mapping of large DNA by chromosome fragmentation. Proc Natl Acad Sci USA 85:6027–6031
Woolford CA, Daniels LB, Park FJ, Jones EW, Van Arsdell JN, Innis MA (1986) The PEP4 gene encodes an aspartyl protease implicated in the posttranslational regulation of Saccharomyces cerevisiae vacuolar hydrolases. Mol Cell Biol 6:2500–2510
Zaret KS, Sherman F (1982) DNA sequence required for efficient transcription termination in yeast. Cell 28:563–573
Author information
Authors and Affiliations
Additional information
Communicated by C.P. Hollenberg
Rights and permissions
About this article
Cite this article
Kaneko, Y., Toh-e, A., Banno, I. et al. Molecular characterization of a specific p-nitrophenylphosphatase gene, PHO13, and its mapping by chromosome fragmentation in Saccbaromyces cerevisiae . Molec. Gen. Genet. 220, 133–139 (1989). https://doi.org/10.1007/BF00260867
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00260867