Abstract
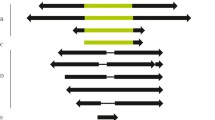
The mobile element ZAM, recently identified in Drosophila melanogaster, is similar in structure and coding potential to vertebrate retroviruses. In this paper, we analyze the insertional and structural polymorphism of this element and show that members of this family appear to have a long evolutionary history in the genome of Drosophila. It is present in all the species of the D. melanogaster subgroup and in more distantly related species like D. takahashii, D. ananassae, or D. virilis but in a lower copy number or with a lower homology. Two categories of strains have been previously identified in D. melanogaster: strains with a high copy number of ZAM and strains with a low copy number. Here, we show that ZAM is at least in a low copy number in each tested strain of the species analyzed. The study of ZAM's genomic distribution by FISH mapping analysis to salivary gland polytene chromosomes or on mitotic chromosomes indicates that most of the insertion sites of ZAM elements are associated with the constitutive heterochromatin regardless of the ZAM copy number. In addition, our results suggest that multiple ZAM elements are present at the insertion sites visualized by in situ experiments.
Similar content being viewed by others
References
Alberola, T.M. & R. De Frutos, 1996. Molecular structure of a Gypsy element of Drosophila subobscura (gypsyDs) constituting a degenerate form of insect retroviruses. Nucleic Acids Res. 24: 914–923.
Bucheton, A., M. Simonelig, C. Vaury & M. Crozatier, 1986. Sequences similar to the transposable element involved in I-R hybrid dysgenesis in D. melanogaster occur in other Drosophila species. Nature 322: 650–652.
Daniels, S.B. & L.D. Strausbaugh, 1986. The distribution of P-element sequences in Drosophila:thewillistoni and saltans species groups. J. Mol. Evol. 17: 138–148.
David, J., 1959. Etude quantitative du dèveloppement de la Drosophile èlevèe en milieu axènique. Bull. Soc. Biol., France & Belgique 93: 472.
De Frutos, R., K.R. Peterson & M.G. Kidwell, 1992. Distribution of Drosophila melanogaster transposable element sequences in species of the obscura group. Chromosoma 101: 293–300.
Gatti, M., S. Bonascossi & S. Pimpinelli, 1994. Looking at Drosophila mitotic chromosomes. Methods in Cell Biology 44: 371–391.
Hochstenbach R., H. Harhangi, K. Schouren, P. Bindels, R. Suijker-buijk & W. Hennig, 1996. Transcription of gypsy elements in a Y-chromosome male fertility gene of Drosophila hydei. Genetics 142: 437–446.
Kim, A.I., C. Terzian, P. Santamaria, A. Pélisson, N. Prud'homme & A. Bucheton, 1994. Retroviruses in invertebrates: the gypsy retro-transposon is apparently an infectious retrovirus of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 91: 1285–1289.
Leblanc, P., S. Desset, B. Dastugue & C. Vaury, 1997. Invertebrate retroviruses: ZAM, a new candidate in Drosophila melanogaster. EMBO J. (in press).
Lemeunier, F., J. David, L. Tsacas & M. Ashburner, 1986. The melanogaster species group, pp. 147–256 in The Genetics and Biology of Drosophila Vol. 3e, edited by M. Ashburner, H.L. Carson & J.N. Thompson. Academic Press: London.
Lohe, A., E. Moriyama, D. Lindholm & D. Hartl, 1995. Horizontal transmission, vertical inactivation, and stochastic loss of Mariner-like transposable elements. Mol. Biol. Evol. 11: 40–50.
Maruyama, K. & D.L. Hartl, 1991. Evidence for interspecific transfer of the transposable element mariner between Drosophila and Zaprionus. J. Mol. Evol. 33: 514–524.
Miklos, G.L.G. & J.N. Cotsell, 1990. Chromosome structure at interfaces between major chromatin types: α-and β-heterochromatin. BioEssays 12: 1–7.
Mizrokhi, L.J. & A.M. Mazo, 1990. Evidence for horizontal transmission of the mobile element jockey between distant Drosophila species. Proc. Natl. Acad. Sci. USA 87: 9216–9220.
Pimpinelli, S. & P. Dimitri, 1989. Cytogenetic analysis of segregation distortion in Drosophila melanogaster: the cytological organization of the Responder (Rsp) locus. Genetics 121: 765–772.
Pimpinelli, S., M. Berloco, L. Fanti, P. Dimitri, S. Bonaccorsi, E. Marchetti, R. Caizzi, C. Caggese & M. Gatti, 1995. Transposable elements are stable structural components of Drosophila melanogaster heterochromatin. Proc. Natl. Acad. Sci. USA 92: 3804–3808.
Sambrook, J., E.F. Fritsch & T. Maniatis, 1989. Molecular cloning: a laboratory manual (2nd Edn.). Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
Stacey, S.N., R.A. Lansman, H.W. Brock & T.A. Grigliatti, 1986. Distribution and conservation of mobile elements in the genus Drosophila. Mol. Biol. Evol. 3: 522–534.
Throckmorton, L.H, 1975. The phylogeny, ecology and geography of Drosophila, pp. 421–469 in Handbook of Genetics vol. 3, edited by R.C. King. Plenum: New York.
Vaury, C., A. A. Bucheton & Pélisson, 1989. The β-heterochromatic sequences flanking the I elements are themselves defective transposable elements. Chromosoma. 98: 215–224.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Baldrich, E., Dimitri, P., Desset, S. et al. Genomic distribution of the retrovirus-like element ZAM in Drosophila. Genetica 100, 131–140 (1997). https://doi.org/10.1023/A:1018317209658
Issue Date:
DOI: https://doi.org/10.1023/A:1018317209658