Abstract
Microbiomes are important to the survival and reproduction of their hosts. Although ecological and evolutionary processes can happen simultaneously in microbiomes, little is known about how microbiome eco-evolutionary dynamics determine host fitness. Here we show, using experimental evolution, that fitness of the aquatic plant Lemna minor is modified by interactions between the microbiome and the evolution of one member, Pseudomonas fluorescens. Microbiome presence promotes P. fluorescens’ rapid evolution to form biofilm, which reciprocally alters the microbiome’s species composition. These eco-evolutionary dynamics modify the host’s multigenerational fitness. The microbiome and non-evolving P. fluorescens together promote host fitness, whereas the microbiome with P. fluorescens that evolves biofilm reduces the beneficial impact on host fitness. Additional experiments suggest that the microbial effect on host fitness may occur through changes in microbiome production of auxin, a plant growth hormone. Our study, therefore, experimentally demonstrates the importance of the eco-evolutionary dynamics in microbiomes for host–microbiome interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the finding of this study are available on Figshare (https://doi.org/10.6084/m9.figshare.13644161). Source data are provided with this paper.
References
Slobodkin, L. B. Growth and regulation of animal populations (Holt, Rinehart and Winston, 1961).
Thompson, J. N. Rapid evolution as an ecological process. Trends Ecol. Evol. 13, 329–332 (1998).
Hendry, A. P. A critique for eco-evolutionary dynamics. Funct. Ecol. 33, 84–94 (2019).
Turcotte, M. M., Reznick, D. N. & Hare, J. D. The impact of rapid evolution on population dynamics in the wild: experimental test of eco-evolutionary dynamics. Ecol. Lett. 14, 1084–1092 (2011).
Hairston, N. G. Jr, Ellner, S. P., Geber, M. A., Yoshida, T. & Fox, J. A. Rapid evolution and the convergence of ecological and evolutionary time. Ecol. Lett. 8, 1114–1127 (2005).
Tan, J., Rattray, J. B., Yang, X. & Jiang, L. Spatial storage effect promotes biodiversity during adaptive radiation. Proc. R. Soc. Lond. B 284, 20170841 (2017).
Hart, S. P., Turcotte, M. M. & Levine, J. M. Effects of rapid evolution on species coexistence. Proc. Natl Acad. Sci. USA 116, 2112–2117 (2019).
Faillace, C. A. & Morin, P. J. Evolution alters the consequences of invasions in experimental communities. Nat. Ecol. Evol. 1, 0013 (2017).
Vanbergen, A. J., Espíndola, A. & Aizen, M. A. Risks to pollinators and pollination from invasive alien species. Nat. Ecol. Evol. 2, 16–25 (2018).
Hendry, A. P. Eco-evolutionary dynamics (Princeton Univ. Press, 2016).
Garud, N. R., Good, B. H., Hallatschek, O. & Pollard, K. S. Evolutionary dynamics of bacteria in the gut microbiome within and across hosts. PLoS Biol. 17, e3000102 (2019).
Zhao, S. et al. Adaptive evolution within gut microbiomes of healthy people. Cell Host Microbe 25, 656–667 (2019).
terHorst, C. P. & Zee, P. C. Eco-evolutionary dynamics in plant–soil feedbacks. Funct. Ecol. 30, 1062–1072 (2016).
Soto, M. J., Domínguez‐Ferreras, A., Pérez‐Mendoza, D., Sanjuán, J. & Olivares, J. Mutualism versus pathogenesis: the give‐and‐take in plant–bacteria interactions. Cell. Microbiol. 11, 381–388 (2009).
Marchetti, M. et al. Experimental evolution of a plant pathogen into a legume symbiont. PLoS Biol. 8, e1000280 (2010).
Saikkonen, K., Wäli, P., Helander, M. & Faeth, S. H. Evolution of endophyte–plant symbioses. Trends Plant Sci. 9, 275–280 (2004).
Reese, A. T. & Dunn, R. R. Drivers of microbiome biodiversity: a review of general rules, feces, and ignorance. mBio 9, e01294-18 (2018).
Miller, E. T., Svanbäck, R. & Bohannan, B. J. Microbiomes as metacommunities: understanding host-associated microbes through metacommunity ecology. Trends Ecol. Evol. 33, 926–935 (2018).
Griffin, E. A. et al. Plant host identity and soil macronutrients explain little variation in sapling endophyte community composition: is disturbance an alternative explanation? J. Ecol. 107, 1876–1889 (2019).
Acosta, K. et al. Duckweed hosts a taxonomically similar bacterial assemblage as the terrestrial leaf microbiome. PLoS ONE 15, e0228560 (2020).
Sandler, G., Bartkowska, M., Agrawal, A. F. & Wright, S. I. Estimation of the SNP mutation rate in two vegetatively propagating species of duckweed. G3 10, 4191–4200 (2020).
Ishizawa, H., Kuroda, M., Morikawa, M. & Ike, M. Evaluation of environmental bacterial communities as a factor affecting the growth of duckweed Lemna minor. Biotechnol. Biofuels 10, 62 (2017).
Zhang, Y. et al. Duckweed (Lemna minor) as a model plant system for the study of human microbial pathogenesis. PLoS ONE 5, e13527 (2010).
Rainey, P. B. & Travisano, M. Adaptive radiation in a heterogeneous environment. Nature 394, 69–72 (1998).
Tan, J., Yang, X., He, Q., Hua, X. & Jiang, L. Earlier parasite arrival reduces the repeatability of host adaptive radiation. ISME J. 14, 2358–2360 (2020).
Tan, J., Yang, X. & Jiang, L. Species ecological similarity modulates the importance of colonization history for adaptive radiation. Evolution 71, 1719–1727 (2017).
Meyer, J. R., Schoustra, S. E., Lachapelle, J. & Kassen, R. Overshooting dynamics in a model adaptive radiation. Proc. R. Soc. Lond. B 278, 392–398 (2011).
Tan, J., Kelly, C. K. & Jiang, L. Temporal niche promotes biodiversity during adaptive radiation. Nat. Commun. 4, 2102 (2013).
Spiers, A. J., Buckling, A. & Rainey, P. B. The causes of Pseudomonas diversity. Microbiology 146, 2345–2350 (2000).
Spiers, A. J., Bohannon, J., Gehrig, S. M. & Rainey, P. B. Biofilm formation at the air–liquid interface by the Pseudomonas fluorescens SBW25 wrinkly spreader requires an acetylated form of cellulose. Mol. Microbiol. 50, 15–27 (2003).
Bantinaki, E. et al. Adaptive divergence in experimental populations of Pseudomonas fluorescens. III. Mutational origins of wrinkly spreader diversity. Genetics 176, 441–453 (2007).
McDonald, M. J., Gehrig, S. M., Meintjes, P. L., Zhang, X.-X. & Rainey, P. B. Adaptive divergence in experimental populations of Pseudomonas fluorescens. IV. Genetic constraints guide evolutionary trajectories in a parallel adaptive radiation. GENETICS 183, 1041–1053 (2009).
Bailey, S. F., Dettman, J. R., Rainey, P. B. & Kassen, R. Competition both drives and impedes diversification in a model adaptive radiation. Proc. R. Soc. Lond. B 280, 20131253 (2013).
Hansen, S. K., Rainey, P. B., Haagensen, J. A. & Molin, S. Evolution of species interactions in a biofilm community. Nature 445, 533–536 (2007).
Flemming, H.-C. et al. Biofilms: an emergent form of bacterial life. Nat. Rev. Microbiol. 14, 563–575 (2016).
Ahmad, F., Ahmad, I. & Khan, M. S. Screening of free-living rhizospheric bacteria for their multiple plant growth promoting activities. Microbiol. Res. 163, 173–181 (2008).
El-Sayed, W. S., Akhkha, A., El-Naggar, M. Y. & Elbadry, M. In vitro antagonistic activity, plant growth promoting traits and phylogenetic affiliation of rhizobacteria associated with wild plants grown in arid soil. Front. Microbiol. 5, 651 (2014).
Gómez, P. & Buckling, A. Real-time microbial adaptive diversification in soil. Ecol. Lett. 16, 650–655 (2013).
Spor, A., Koren, O. & Ley, R. Unravelling the effects of the environment and host genotype on the gut microbiome. Nat. Rev. Microbiol. 9, 279–290 (2011).
Walters, W. A. et al. Large-scale replicated field study of maize rhizosphere identifies heritable microbes. Proc. Natl Acad. Sci. USA 115, 7368–7373 (2018).
Veach, A. M. et al. Rhizosphere microbiomes diverge among Populus trichocarpa plant-host genotypes and chemotypes, but it depends on soil origin. Microbiome 7, 76 (2019).
Lennon, J. T. & Martiny, J. B. Rapid evolution buffers ecosystem impacts of viruses in a microbial food web. Ecol. Lett. 11, 1178–1188 (2008).
Pantel, J. H., Duvivier, C. & Meester, L. D. Rapid local adaptation mediates zooplankton community assembly in experimental mesocosms. Ecol. Lett. 18, 992–1000 (2015).
Faillace, C. A. & Morin, P. J. Evolution alters post-invasion temporal dynamics in experimental communities. J. Anim. Ecol. 89, 285–298 (2020).
Omilian, A. R., Cristescu, M. E. A., Dudycha, J. L. & Lynch, M. Ameiotic recombination in asexual lineages of Daphnia. Proc. Natl Acad. Sci. USA 103, 18638–18643 (2006).
Mao, Y., Botella, J. R., Liu, Y. & Zhu, J.-K. Gene editing in plants: progress and challenges. Natl Sci. Rev. 6, 421–437 (2019).
Horvath, P. & Barrangou, R. CRISPR/Cas, the immune system of Bacteria and Archaea. Science 327, 167–170 (2010).
Yang, L. et al. Promotion of plant growth and in situ degradation of phenol by an engineered Pseudomonas fluorescens strain in different contaminated environments. Soil Biol. Biochem. 43, 915–922 (2011).
Zabłocka-Godlewska, E., Przystaś, W. & Grabińska-Sota, E. Decolourization of diazo Evans blue by two strains of Pseudomonas fluorescens isolated from different wastewater treatment plants. Water Air Soil Pollut. 223, 5259–5266 (2012).
Paulsen, I. T. et al. Complete genome sequence of the plant commensal Pseudomonas fluorescens Pf-5. Nat. Biotechnol. 23, 873–878 (2005).
Rainey, P. B. Adaptation of Pseudomonas fluorescens to the plant rhizosphere. Environ. Microbiol. 1, 243–257 (1999).
Gilbert, S. et al. Bacterial production of indole related compounds reveals their role in association between duckweeds and endophytes. Front. Chem. 6, 265 (2018).
Bailey, M. J., Lilley, A. K., Thompson, I. P., Rainey, P. B. & Ellis, R. J. Site directed chromosomal marking of a fluorescent pseudomonad isolated from the phytosphere of sugar beet; stability and potential for marker gene transfer. Mol. Ecol. 4, 755–764 (1995).
Spiers, A. J. & Rainey, P. B. The Pseudomonas fluorescens SBW25 wrinkly spreader biofilm requires attachment factor, cellulose fibre and LPS interactions to maintain strength and integrity. Microbiology 151, 2829–2839 (2005).
Lind, P. A., Libby, E., Herzog, J. & Rainey, P. B. Predicting mutational routes to new adaptive phenotypes. eLife 8, e38822 (2019).
O’Brien, P. A., Webster, N. S., Miller, D. J. & Bourne, D. G. Host–microbe coevolution: applying evidence from model systems to complex marine invertebrate holobionts. mBio 10, e02241-18 (2019).
Theis, K. R. et al. Getting the hologenome concept right: an eco-evolutionary framework for hosts and their microbiomes. mSystems 1, e00028-16 (2016).
Landolt, E. Biosystematic Investigations in the Family of Duckweeds (Lemnaceae), Volume 2. The Family of Lemnaceae, A Monographic Study, Volume 1 (Geobotanical Institute, ETH Zurich, 1986).
Ziegler, P., Sree, K. S. & Appenroth, K.-J. Duckweeds for water remediation and toxicity testing. Toxicol. Environ. Chem. 98, 1127–1154 (2016).
Acknowledgements
We thank P. Rainey for providing us with the two P. fluorescens strains and J. Armstrong, D. Conover and A. Morris for collecting and genotyping L. minor. We thank J. Everett, L. Leak, S. Subramanian, E. Elliott and R. Dabundo for assistance with the experiment and K. Kohl, L. Rzodkiewicz, E. Gluck-Thaler, N. Wei and C. Wood for comments that improved the manuscript. This project is supported by a British Ecological Society Research Grant (LRB17/1023).
Author information
Authors and Affiliations
Contributions
J.T. and M.M.T. conceived the idea and designed the study. J.T. and J.E.K. performed the experiments. J.T., J.E.K. and M.M.T. analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Ecology & Evolution thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Results of the mutual invasion experiments between the various P. fluorescens lineages.
The goal of these experiments is to test whether the ancestral free-living SBW25 (smooth morph, SM) is ecologically similar to the isogenic PBR716 (SM) with the three operon knock-outs. We also quantified the ecological differences between wrinkly spreader (WS) and SM of SBW25 for comparison. Between SM of PBR716 and SM of SBW25, we first quantified their growth in monoculture (rmono) in aqueous microcosms. We set the initial abundance at ~106 CFU (0.01x of carrying capacity). We quantified bacterial abundance after 24 h of growth and calculated rmono as ln (abundance). We quantified their invading growth (rinvading) in a culture of the other strain that had been already growing to high abundance for 24 h. The growth of the invading strain was calculated based on its final abundance after 24 h static incubation. Please note that SBW25 strain we used carried a neutral lacZ marker while PBR716 did not. Therefore, we distinguished the SM of the two strains based on their colony color on with X-gal (SBW25, blue; PBR716, white). All treatments were replicated four times. We quantified the competitive response (S) of one strain to the other as 1- rinvading/rmono. S close to 1 indicates stronger interaction and potentially higher ecological similarity between two strains, while that close to 0 indicates weaker interaction and lower ecological similarity. We showed that higher competitive responses between SM of PBR716 and SM of SBW25 than those between SM and WS of SBW25. Values are means (± 1 s.e.; n = 4). Treatments sharing the same letter are not statistically different.
Extended Data Fig. 2 The abundance of the microbiome species subject to three P. fluorescens treatments (No Pf, non-evolving PBR716, and evolving SBW25).
The microbiome contained Bacillus pumilus (BP), Agrobacterium tumefaciens (AT), Sphingomonas elodea (SE), Rhizobium rosettiformans (RR), Chryseobacterium hispalense (CH), Duganella radices (DR), Variovorax paradoxus (VP), and Flavobacterium buctense (FB) (n for each species = 12). Bacillus aryabhattai was isolated from L. minor epiphyte, but appeared only in the medium, and was therefore not shown here. Boxes show medians and interquartile ranges with whiskers for 10th and 90th percentiles. Stars indicate that the abundance of A. tumefaciens and R. rosettiformans was significantly influenced by P. fluorescens presence or evolution.
Extended Data Fig. 3 The abundance of L. minor in each L. minor genotype treatment.
Boxes show medians and interquartile ranges with whiskers for 10th and 90th percentiles (n = 4). Treatments sharing the same letter are not statistically different (P > 0.05). Pf stands for Pseudomonas fluorescens. L. minor was genotyped with two microsatellite primers R5C (F: TGATGCCAGTAGATCCGGC R: ACGCCTGAACACGATTGATG) and R15B (F: GTGACAGCGTATCCTTGTGC R: TCAGCGGCAAGATCATCAAG).
Extended Data Fig. 4 The concentration of available phosphorus.
Values are means (± 1 s.e.; n = 3). Pf stands for Pseudomonas fluorescens. Treatments sharing the same letter are not statistically different (P > 0.05).
Extended Data Fig. 5 The correlation between auxin production and L. minor fitness.
After the main experiment ended, we plated the culture from each microcosm on agar plates to quantify the abundance of bacterial species/genotypes. We isolated each present species/genotype and re-assembled the microbiome in a new microcosm with L. minor. The microbiome was incubated for a week to allow them to propagate and equilibrate. Then, we added them into a new microcosm without L. minor to quantify auxin concentration after 24 h. The average level of auxin production of the six experimental treatments (microbiome presence/absence × P. fluorescens treatments) were calculated and presented in Fig. 4. Here, we plot the number of L. minor individuals of each treatment (data collected from the main eperiment, y-axis data, ln-transformed before the analysis) against auxin production (x-axis; N = 72). We implemented both a linear (y=a – bx) and asymptotic (y=a – [a – b] exp [–cx]) model in R. The asymptotic model had a lower AIC value (asymptotic 529.410, linear 557.702; P values for both models is smaller than 0.001) and thus better predict the relationship between auxin production and L. minor fitness.
Supplementary information
Supplementary Information
Supplementary Tables 1–6.
Source data
Source Data Fig. 1
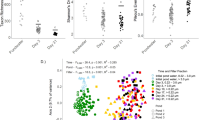
Source data for wrinkly spreader proportion and P. fluorescens abundance.
Source Data Fig. 2
Source data for NMDS.
Source Data Fig. 3
Source data for L. minor fitness.
Source Data Fig. 4
Source data for microbiome-auxin-production assay and auxin-addition experiment.
Source Data Extended Data Fig. 1
Source data for SBW25 and PBR716 competitive responses.
Source Data Extended Data Fig. 2
Source data for bacterial abundance.
Source Data Extended Data Fig. 3
Source data for the fitness of the three L. minor genotypes.
Source Data Extended Data Fig. 4
Source data for phosphorus concentration.
Source Data Extended Data Fig. 5
Source data for the auxin–L. minor fitness correlation.
Rights and permissions
About this article
Cite this article
Tan, J., Kerstetter, J.E. & Turcotte, M.M. Eco-evolutionary interaction between microbiome presence and rapid biofilm evolution determines plant host fitness. Nat Ecol Evol 5, 670–676 (2021). https://doi.org/10.1038/s41559-021-01406-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-021-01406-2
This article is cited by
-
Eco-evolutionary interactions and host fitness
Nature Reviews Microbiology (2021)