Abstract
Aims
Hotspots of enzyme activity in soil strongly depend on carbon inputs such as rhizodeposits and root detritus. In this study, we compare the effect of living and dead Lupinus polyphyllus L. roots on the small-scale distribution of cellulase, chitinase and phosphatase activity in soil.
Methods
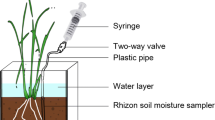
Soil zymography, a novel in situ method, was used to analyze extracellular cellulase, chitinase and phosphatase activity in the presence of i. living L. polyphyllus roots prior to shoot cutting and ii. dead/dying roots 10, 20 and 30 days after shoot cutting.
Results
After shoot cutting, cellulase and chitinase activities increased and were highest at the root tips. The areas of high cellulase and phosphatase activity extend up to 55 mm away from the root. Moreover, we observed microhotspots of cellulose, chitinase, and phosphatase activity up to 60 mm away from the next living root. The number and activity of microhotspots of chitinase activity was maximal 10 days after shoot cutting.
Conclusions
The study showed that young root detritus stimulates enzyme activities stronger than living roots. Soil zymography allowed identification of microhotspots of enzyme activity up to several cm away from living and dying roots, which most likely were caused by arbuscular mycorrhizal fungi.







Similar content being viewed by others
References
Allison SD (2005) Cheaters, diffusion and nutrients constrain decomposition by microbial enzymes in spatially structured environments. Ecol Lett 8:626–635
Bastian F, Bouziri L, Nicolardot B, Ranjard L (2009) Impact of wheat straw decomposition on successional patterns of soil microbial community structure. Soil Biol Biochem 41:262–275
Blagodatskaya E, Kuzyakov Y (2013) Active microorganisms in soil: critical review of estimation criteria and approaches. Soil Biol Biochem 67:197–211
Butenschoen O, Marhan S, Scheu S (2008) Response of soil microorganisms and endogeic earthworms to cutting of grassland plants in a laboratory experiment. Appl Soil Ecol 38:152–160
Davey ME, O’toole GA (2000) Microbial biofilms: from ecology to molecular genetics. Microbiol Mol Biol Rev 64:847–867
De Nobili M, Contin M, Mondini C, Brookes PC (2001) Soil microbial biomass is triggered into activity by trace amounts of substrate. Soil Biol Biochem 33:1163–1170
Dong S, Brooks D, Jonesa MD, Grayston SJ (2007) A method for linking in situ activities of hydrolytic enzymes to associated organisms in forest soils. Soil Biol Biochem 39:2414–2435
Ekschmitt K, Liu MQ, Vetter S, Fox O, Wolters V (2005) Strategies used by soil biota to overcome soil organic matter stability–why is dead organic matter left over in the soil? Geoderma 128:167–176
Gill RA, Jackson RB (2000) Global patterns of root turnover for terrestrial ecosystems. New Phytol 147:13–31
Guitian R, Bardgett RD (2000) Plant and soil microbial responses to defoliation in temperate semi-natural grassland. Plant Soil 220:271–277
Henriksen TM, Breland TA (2002) Carbon mineralization, fungal and bacterial growth, and enzyme activities as affected by contact between crop residues and soil. Biology and Fertility of Soils 35:41–48.
Hinsinger P, Bengough A, Vetterlein D, Young I (2009) Rhizosphere: biophysics, biogeochemistry and ecological relevance. Plant Soil 321:117–152
Högberg P, Read DJ (2006) Towards a more plant physiological perspective on soil ecology. Trends Ecol Evol 21:548–554
Joergensen RG, Brookes PC, Jenkinson DS (1990) Survival of the soil microbial biomass at elevated temperatures. Soil Biol Biochem 22:1129–1136
Jones DL, Nguyen C, Finlay RD (2009) Carbon flow in the rhizosphere: carbon trading at the soil–root interface. Plant Soil 321:5–33
Kandeler E, Luxhøi J, Tscherko D, Magid J (1999) Xylanase, invertase and protease at the soil–litter interface of a loamy sand. Soil Biol Biochem 31:1171–1179
Kandeler E, Marschner P, Tscherko D, Gahoonia TS, Nielsen NE, (2002) Microbial community composition and functional diversity in the rhizosphere of maize. Plant Soil 238:301–312
Kuzyakov Y (2002) Factors affecting rhizosphere priming effects. J Plant Nutr Soil Sci 165:382–396
Kuzyakov Y, Cheng W (2001) Photosynthesis controls of rhizosphere respiration and organic matter decomposition. Soil Biol Biochem 33:1915–1925
Leake JR, Donnelly DP, Saunders EM, Boddy L, Read DJ (2001) Rates and quantities of carbon flux to ectomycorrhizal mycelium following 14C pulse labeling of Pinus sylvestris seedlings: effects of litter patches and interaction with a wood-decomposer fungus. Tree Physiol 21:71–82
Luster J, Göttlein A, Nowack B, Sarret G (2009) Sampling, defining, characterising and modeling the rhizosphere—the soil science tool box. Plant Soil 321:457–482
Marschner P, Marhan S, Kandeler E (2012) Microscale distribution and function of soil microorganisms in the interface between rhizosphere and detritusphere. Soil Biol Biochem 49:174–183
Nguyen C (2003) Rhizodeposition of organic C by plants: mechanisms and controls. Agron Sci Prod Veg Environ 23:375–396
Nunan N, Wu K, Young MN, Crawford JW, Ritz K (2002) In situ spatial patterns of soil bacterial populations, mapped at multiple scales, in an arable soil. Microb Ecol 44:296–305
Parkin TB (1987) Soil microsites as a source of denitrification variability. Soil Sci Soc Am J 51:1194–1199
Poll C, Ingwersen J, Shootmer M, Gerzabek MH, Kandeler E (2006) Mechanisms of solute transport affect small–scale abundance and function of soil microorganisms in the detritusphere. Eur J Soil Sci 57:583–595
Poll C, Marhan S, Ingwersen J, Kandeler E (2008) Dynamics of litter carbon turnover and microbial abundance in a rye detritusphere. Soil Biol Biochem 40:1306–1321
Rasse DP, Rumpel C, Dignac MF (2005) Is soil carbon mostly root carbon? Mechanisms for a specific stabilisation. Plant Soil 269:341–356
R Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/
Schimel JP, Weintraub MN (2003) The implications of exoenzyme activity on microbial carbon and nitrogen limitation in soil: a theoretical model. Soil Biol Biochem 35:549–563
Spohn M, Kuzyakov Y (2013) Distribution of microbial- and root-derived phosphatase activities in the rhizosphere depending on P availability and C allocation–coupling soil zymography with 14C imaging. Soil Biol Biochem 67:106–113
Spohn M, Carminati A, Kuzyakov Y (2013) Soil zymography—a novel in situ method for mapping distribution of enzyme activity in soil. Soil Biol Biochem 58:275–280
Tarafdar JC, Jungk A (1987) Phosphatase activity in the rhizosphere and its relation to the depletion of soil organic phosphorus. Biol Fertil Soils 3:199–204
Vance ED, Chapin FS III (2001) Substrate limitations to microbial activity in taiga forest floors. Soil Biol Biochem 33:173–188
Watt M, Wendy KS, Passioura JB (2006) Rates of root and organism growth, soil conditions, and temporal and spatial development of the rhizosphere. Ann Bot 97:839–855
Westover KM, Kennedy AC, Kelley SE (1997) Patterns of rhizosphere microbial community structure associated with co-occurring plant species. J Ecol 85:863–873
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Stefano Manzoni.
Rights and permissions
About this article
Cite this article
Spohn, M., Kuzyakov, Y. Spatial and temporal dynamics of hotspots of enzyme activity in soil as affected by living and dead roots—a soil zymography analysis. Plant Soil 379, 67–77 (2014). https://doi.org/10.1007/s11104-014-2041-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-014-2041-9