Abstract
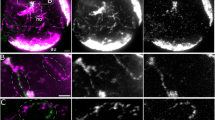
The Y loops of Drosophila spermatocytes are formed by the expression of huge individual transcription units on the Y chromosome and their large size provides a unique system for the investigation of the organisation of transcription in intact nuclei. By labelling ribonucleo-protein (RNP) components, the loop chromatin and nascent transcripts in Y loop C, we reveal a highly structured organisation of RNP domains associated with nascent transcripts. We distinguish two types of RNP domain, a proximal domain that runs alongside the chromatin of loop C and a distal RNP domain that wraps around the proximal domain and the loop chromatin. The proximal domain is marked by the Pasilla protein, and separate distal subdomains are marked by the S5 antigen and Boule. We discuss the implications of this highly structured co-transcriptional architecture for the organisation of the process of transcription.





Similar content being viewed by others
References
Abramoff MD, Magelhaes PJ, Ram SJ (2004) Image processing with ImageJ. Biophotonics Int 11(7):36–42
Alonso-Gonzalez L, Dominguez A, Albornoz J (2003) Structural heterogeneity and genomic distribution of Drosophila melanogaster LTR-retrotransposons. Mol Biol Evol 20(3):401–409
Bonaccorsi S, Lohe A (1991) Fine mapping of satellite DNA sequences along the Y chromosome of Drosophila melanogaster: relationships between satellite sequences and fertility factors. Genetics 129(1):177–189
Bonaccorsi S, Pisano C, Puoti F, Gatti M (1988) Y chromosome loops in Drosophila melanogaster. Genetics 120(4):1015–1034
Bonaccorsi S, Gatti M, Pisano C, Lohe A (1990) Transcription of a satellite DNA on two Y chromosome loops of Drosophila melanogaster. Chromosoma 99(4):260–266
Borie N, Maisonhaute C, Sarrazin S, Loevenbruck C, Biemont C (2002) Tissue-specificity of 412 retrotransposon expression in Drosophila simulans and D. melanogaster. Heredity 89(4):247–252
Brookman JJ, Toosy AT, Shashidhara LS, White RA (1992) The 412 retrotransposon and the development of gonadal mesoderm in Drosophila. Development 116(4):1185–1192
Callan HG (1986) Lampbrush chromosomes. Mol Biol Biochem Biophys 36:1–252
Carvalho AB, Lazzaro BP, Clark AG (2000) Y chromosomal fertility factors kl-2 and kl-3 of Drosophila melanogaster encode dynein heavy chain polypeptides. Proc Natl Acad Sci USA 97(24):13239–13244
Carvalho AB, Dobo BA, Vibranovski MD, Clark AG (2001) Identification of five new genes on the Y chromosome of Drosophila melanogaster. Proc Natl Acad Sci USA 98(23):13225–13230
Cheng MH, Maines JZ, Wasserman SA (1998) Biphasic subcellular localization of the DAZL-related protein boule in Drosophila spermatogenesis. Dev Biol 204(2):567–576
Clarkson M, Saint R (1999) A His2AvDGFP fusion gene complements a lethal His2AvD mutant allele and provides an in vivo marker for Drosophila chromosome behavior. DNA Cell Biol 18(6):457–462
Cook PR (1999) The organization of replication and transcription. Science 284(5421):1790–1795
Cook PR (2009) A model for all genomes: the role of transcription factories. J Mol Biol 395:1–10
Gepner J, Hays TS (1993) A fertility region on the Y chromosome of Drosophila melanogaster encodes a dynein microtubule motor. Proc Natl Acad Sci USA 90(23):11132–11136
Glatzer KH (1984) Preservation of nuclear RNP antigens in male germ cell development of Drosophila hydei. Mol Gen Genet 196:236–243
Glatzer KH, Meyer GF (1981) Morphological aspects of the genetic activity in primary spermatocyte nuclei of Drosophila hydei. Biol Cell 41:165–172
Grond CJ, Siegmund I, Hennig W (1983) Visualization of a lampbrush loop-forming fertility gene in Drosophila hydei. Chromosoma 88:50–56
Hackstein JH, Hochstenbach R (1995) The elusive fertility genes of Drosophila: the ultimate haven for selfish genetic elements. Trends Genet 11(5):195–200
Jensen KB, Dredge BK, Stefani G, Zhong R, Buckanovich RJ, Okano HJ, Yang YY, Darnell RB (2000) Nova-1 regulates neuron-specific alternative splicing and is essential for neuronal viability. Neuron 25(2):359–371
Kornberg RD (1974) Chromatin structure: a repeating unit of histones and DNA. Science 184(139):868–871
Kosman D, Mizutani CM, Lemons D, Cox WG, McGinnis W, Bier E (2004) Multiplex detection of RNA expression in Drosophila embryos. Science 305(5685):846
Lifschytz E, Hareven D, Azriel A, Brodsly H (1983) DNA clones and RNA transcripts of four lampbrush loops from the Y chromosome of Drosophila hydei. Cell 32(1):191–199
Liu LF, Wang JC (1987) Supercoiling of the DNA template during transcription. Proc Natl Acad Sci USA 84(20):7024–7027
Lu AQ, Beckingham K (2000) Androcam, a Drosophila calmodulin-related protein, is expressed specifically in the testis and decorates loop kl-3 of the Y chromosome. Mech Dev 94(1–2):171–181
Park JW, Parisky K, Celotto AM, Reenan RA, Graveley BR (2004) Identification of alternative splicing regulators by RNA interference in Drosophila. Proc Natl Acad Sci USA 101(45):15974–15979
Perales R, Bentley D (2009) “Cotranscriptionality”: the transcription elongation complex as a nexus for nuclear transactions. Mol Cell 36(2):178–191
Reugels AM, Kurek R, Lammermann U, Bunemann H (2000) Mega-introns in the dynein gene DhDhc7(Y) on the heterochromatic Y chromosome give rise to the giant threads loops in primary spermatocytes of Drosophila hydei. Genetics 154(2):759–769
Risau W, Symmons P, Saumweber H, Frasch M (1983) Nonpackaging and packaging proteins of hnRNA in Drosophila melanogaster. Cell 33(2):529–541
Ryder E, Spriggs H, Drummond E, St Johnston D, Russell S (2009) The Flannotator—a gene and protein expression annotation tool for Drosophila melanogaster. Bioinformatics 25(4):548–549
Saumweber H, Symmons P, Kabisch R, Will H, Bonhoeffer F (1980) Monoclonal antibodies against chromosomal proteins of Drosophila melanogaster: establishment of antibody producing cell lines and partial characterization of corresponding antigens. Chromosoma 80(3):253–275
Sutherland H, Bickmore WA (2009) Transcription factories: gene expression in unions? Nat Rev Genet 10(7):457–466
Thummel CS, Burtis KC, Hogness DS (1990) Spatial and temporal patterns of E74 transcription during Drosophila development. Cell 61(1):101–111
Acknowledgements
We are grateful to Kate Beckingham, Harald Saumweber and Steve Wasserman for antibodies. We thank Sarah Bray, Dennis Bray, Alena Krejci and Stephen Young for their comments on the manuscript. The project was funded by the Biotechnology and Biological Sciences Research Council, UK.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Pimpinelli
Rights and permissions
About this article
Cite this article
Redhouse, J.L., Mozziconacci, J. & White, R.A.H. Co-transcriptional architecture in a Y loop in Drosophila melanogaster . Chromosoma 120, 399–407 (2011). https://doi.org/10.1007/s00412-011-0321-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-011-0321-1