Abstract
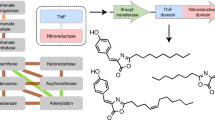
Experimental validation of enzyme function is crucial for genome interpretation, but it remains challenging because it cannot be scaled up to accommodate the constant accumulation of genome sequences. We tackled this issue for the MetA and MetX enzyme families, phylogenetically unrelated families of acyl-L-homoserine transferases involved in L-methionine biosynthesis. Members of these families are prone to incorrect annotation because MetX and MetA enzymes are assumed to always use acetyl-CoA and succinyl-CoA, respectively. We determined the enzymatic activities of 100 enzymes from diverse species, and interpreted the results by structural classification of active sites based on protein structure modeling. We predict that >60% of the 10,000 sequences from these families currently present in databases are incorrectly annotated, and suggest that acetyl-CoA was originally the sole substrate of these isofunctional enzymes, which evolved to use exclusively succinyl-CoA in the most recent bacteria. We also uncovered a divergent subgroup of MetX enzymes in fungi that participate only in L-cysteine biosynthesis as O-succinyl-L-serine transferases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
10 July 2017
In the version of this article initially published, a sentence in the abstract, "Members of these families are prone to incorrect annotation because MetA and MetX enzymes are assumed to always use acetyl-CoA and succinyl-CoA, respectively," had the order of enzymes MetA and MetX reversed. The sentence should read, "Members of these families are prone to incorrect annotation because MetX and MetA enzymes are assumed to always use acetyl-CoA and succinyl-CoA, respectively." The error has been corrected in the HTML and PDF versions of the article.
References
Galperin, M.Y. & Koonin, E.V. From complete genome sequence to 'complete' understanding? Trends Biotechnol. 28, 398–406 (2010).
Hanson, A.D., Pribat, A., Waller, J.C. & de Crécy-Lagard, V. 'Unknown' proteins and 'orphan' enzymes: the missing half of the engineering parts list—and how to find it. Biochem. J. 425, 1–11 (2009).
Schnoes, A.M., Brown, S.D., Dodevski, I. & Babbitt, P.C. Annotation error in public databases: misannotation of molecular function in enzyme superfamilies. PLoS Comput. Biol. 5, e1000605 (2009).
Soskine, M. & Tawfik, D.S. Mutational effects and the evolution of new protein functions. Nat. Rev. Genet. 11, 572–582 (2010).
Bastard, K. et al. Revealing the hidden functional diversity of an enzyme family. Nat. Chem. Biol. 10, 42–49 (2014).
de Crecy-Lagard, V. Quality annotations, a key frontier in the microbial sciences. Microbe 11, 303–310 (2016).
Hsiao, T.L., Revelles, O., Chen, L., Sauer, U. & Vitkup, D. Automatic policing of biochemical annotations using genomic correlations. Nat. Chem. Biol. 6, 34–40 (2010).
Anton, B.P., Kasif, S., Roberts, R.J. & Steffen, M. Objective: biochemical function. Front. Genet. 5, 210 (2014).
Gerlt, J.A. et al. The Enzyme Function Initiative. Biochemistry 50, 9950–9962 (2011).
Zhang, X. et al. Assignment of function to a domain of unknown function: DUF1537 is a new kinase family in catabolic pathways for acid sugars. Proc. Natl. Acad. Sci. USA 113, E4161–E4169 (2016).
Ferla, M.P. & Patrick, W.M. Bacterial methionine biosynthesis. Microbiology 160, 1571–1584 (2014).
Gophna, U., Bapteste, E., Doolittle, W.F., Biran, D. & Ron, E.Z. Evolutionary plasticity of methionine biosynthesis. Gene 355, 48–57 (2005).
de Berardinis, V. et al. A complete collection of single-gene deletion mutants of Acinetobacter baylyi ADP1. Mol. Syst. Biol. 4, 174 (2008).
Foglino, M., Borne, F., Bally, M., Ball, G. & Patte, J.C. A direct sulfhydrylation pathway is used for methionine biosynthesis in Pseudomonas aeruginosa. Microbiology 141, 431–439 (1995).
Rowbury, R.J. The accumulation of O-succinylhomoserine by Escherichia coli and Salmonella typhimurium. J. Gen. Microbiol. 37, 171–180 (1964).
Rowbury, R.J. & Woods, D.D. O-succinylhomoserine as an intermediate in the synthesis of cystathionine by Escherichia coli. J. Gen. Microbiol. 36, 341–358 (1964).
Brush, A. & Paulus, H. The enzymic formation of O-acetylhomoserine in Bacillus subtilis and its regulation by methionine and S-adenosylmethionine. Biochem. Biophys. Res. Commun. 45, 735–741 (1971).
Goudarzi, M. & Born, T.L. Purification and characterization of Thermotoga maritima homoserine transsuccinylase indicates it is a transacetylase. Extremophiles 10, 469–478 (2006).
Rotem, O., Biran, D. & Ron, E.Z. Methionine biosynthesis in Agrobacterium tumefaciens: study of the first enzyme. Res. Microbiol. 164, 12–16 (2013).
Wyman, A. & Paulus, H. Purification and properties of homoserine transacetylase from Bacillus polymyxa. J. Biol. Chem. 250, 3897–3903 (1975).
Ziegler, K., Yusupov, M., Bishop, B. & Born, T.L. Substrate analysis of homoserine acyltransferase from Bacillus cereus. Biochem. Biophys. Res. Commun. 361, 510–515 (2007).
Zubieta, C., Arkus, K.A., Cahoon, R.E. & Jez, J.M. A single amino acid change is responsible for evolution of acyltransferase specificity in bacterial methionine biosynthesis. J. Biol. Chem. 283, 7561–7567 (2008).
Omelchenko, M.V., Galperin, M.Y., Wolf, Y.I. & Koonin, E.V. Non-homologous isofunctional enzymes: a systematic analysis of alternative solutions in enzyme evolution. Biol. Direct 5, 31 (2010).
Hitchcock, D.S. et al. Structure-guided discovery of new deaminase enzymes. J. Am. Chem. Soc. 135, 13927–13933 (2013).
Lukk, T. et al. Homology models guide discovery of diverse enzyme specificities among dipeptide epimerases in the enolase superfamily. Proc. Natl. Acad. Sci. USA 109, 4122–4127 (2012).
de Melo-Minardi, R.C., Bastard, K. & Artiguenave, F. Identification of subfamily-specific sites based on active sites modeling and clustering. Bioinformatics 26, 3075–3082 (2010).
Picardeau, M., Bauby, H. & Saint Girons, I. Genetic evidence for the existence of two pathways for the biosynthesis of methionine in the Leptospira spp. FEMS Microbiol. Lett. 225, 257–262 (2003).
Zhang, Z., Lindstam, M., Unge, J., Peterson, C. & Lu, G. Potential for dramatic improvement in sequence alignment against structures of remote homologous proteins by extracting structural information from multiple structure alignment. J. Mol. Biol. 332, 127–142 (2003).
Hijikata, A., Yura, K., Noguti, T. & Go, M. Revisiting gap locations in amino acid sequence alignments and a proposal for a method to improve them by introducing solvent accessibility. Proteins 79, 1868–1877 (2011).
Centeno, N.B., Planas-Iglesias, J. & Oliva, B. Comparative modelling of protein structure and its impact on microbial cell factories. Microb. Cell Fact. 4, 20 (2005).
Mirza, I.A., Nazi, I., Korczynska, M., Wright, G.D. & Berghuis, A.M. Crystal structure of homoserine transacetylase from Haemophilus influenzae reveals a new family of α/β-hydrolases. Biochemistry 44, 15768–15773 (2005).
Zubieta, C. et al. Crystal structure of homoserine O-succinyltransferase from Bacillus cereus at 2.4 Å resolution. Proteins 68, 999–1005 (2007).
Coe, D.M. & Viola, R.E. Assessing the roles of essential functional groups in the mechanism of homoserine succinyltransferase. Arch. Biochem. Biophys. 461, 211–218 (2007).
Yamagata, S., D'Andrea, R.J., Fujisaki, S., Isaji, M. & Nakamura, K. Cloning and bacterial expression of the CYS3 gene encoding cystathionine γ-lyase of Saccharomyces cerevisiae and the physicochemical and enzymatic properties of the protein. J. Bacteriol. 175, 4800–4808 (1993).
Grynberg, M., Topczewski, J., Godzik, A. & Paszewski, A. The Aspergillus nidulans cysA gene encodes a novel type of serine O-acetyltransferase which is homologous to homoserine O-acetyltransferases. Microbiology 146, 2695–2703 (2000).
Ma, Y. et al. Six new amino acid-auxotrophic markers for targeted gene integration and disruption in fission yeast. Curr. Genet. 52, 97–105 (2007).
Sohn, M.J. et al. Novel cysteine-centered sulfur metabolic pathway in the thermotolerant methylotrophic yeast Hansenula polymorpha. PLoS One 9, e100725 (2014).
Oda, K., Matoba, Y., Kumagai, T., Noda, M. & Sugiyama, M. Crystallographic study to determine the substrate specificity of an L-serine-acetylating enzyme found in the D-cycloserine biosynthetic pathway. J. Bacteriol. 195, 1741–1749 (2013).
Bogicevic, B. et al. Cysteine biosynthesis in Lactobacillus casei: identification and characterization of a serine acetyltransferase. FEMS Microbiol. Lett. 363, fnw012 (2016).
Pedruzzi, I. et al. HAMAP in 2015: updates to the protein family classification and annotation system. Nucleic Acids Res. 43, D1064–D1070 (2015).
Gao, B., Mohan, R. & Gupta, R.S. Phylogenomics and protein signatures elucidating the evolutionary relationships among the Gammaproteobacteria. Int. J. Syst. Evol. Microbiol. 59, 234–247 (2009).
Gupta, R.S. Origin of diderm (Gram-negative) bacteria: antibiotic selection pressure rather than endosymbiosis likely led to the evolution of bacterial cells with two membranes. Antonie van Leeuwenhoek 100, 171–182 (2011).
Jensen, L.J., Ussery, D.W. & Brunak, S. Functionality of system components: conservation of protein function in protein feature space. Genome Res. 13, 2444–2449 (2003).
Khersonsky, O., Roodveldt, C. & Tawfik, D.S. Enzyme promiscuity: evolutionary and mechanistic aspects. Curr. Opin. Chem. Biol. 10, 498–508 (2006).
Furnham, N., Dawson, N.L., Rahman, S.A., Thornton, J.M. & Orengo, C.A. Large-scale analysis exploring evolution of catalytic machineries and mechanisms in enzyme superfamilies. J. Mol. Biol. 428, 253–267 (2016).
Kanzaki, H., Kobayashi, M., Nagasawa, T. & Yamada, H. Distribution of two kinds of cystathionine γ-synthase in various bacteria. FEMS Microbiol. Lett. 33, 65–68 (1986).
Furnham, N., Garavelli, J.S., Apweiler, R. & Thornton, J.M. Missing in action: enzyme functional annotations in biological databases. Nat. Chem. Biol. 5, 521–525 (2009).
Huerta-Cepas, J., Dopazo, J. & Gabaldón, T. ETE: a Python environment for tree exploration. BMC Bioinformatics 11, 24 (2010).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Asnicar, F., Weingart, G., Tickle, T.L., Huttenhower, C. & Segata, N. Compact graphical representation of phylogenetic data and metadata with GraPhlAn. PeerJ 3, e1029 (2015).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Kreimeyer, A. et al. Identification of the last unknown genes in the fermentation pathway of lysine. J. Biol. Chem. 282, 7191–7197 (2007).
Stuani, L. et al. Novel metabolic features in Acinetobacter baylyi ADP1 revealed by a multiomics approach. Metabolomics 10, 1223–1238 (2014).
Sakai, A. et al. Evolution of enzymatic activities in the enolase superfamily: N-succinylamino acid racemase and a new pathway for the irreversible conversion of D- to L-amino acids. Biochemistry 45, 4455–4462 (2006).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Trott, O. & Olson, A.J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Acknowledgements
We are grateful to L. Stuani, L. Grenot, C. Gazaille, C. Richer, C. Pellé and P. Sirvain for excellent technical assistance. We thank G. Cohen, M. Bouzon-Bloch, M. Stam, A. Calteau, B. Viart and M. Sorokina for helpful discussion on the manuscript. We are indebted to P. Bowe for improvements to the manuscript. We are grateful to E. Coudert and C. Rivoire (the Swiss-Prot Group at the SIB Swiss Institute of Bioinformatics), who updated the HAMAP rules. This work was supported by Commissariat à l'énergie atomique et aux énergies alternatives (CEA), the CNRS and the University of Evry Val d'Essonne.
Author information
Authors and Affiliations
Contributions
V.d.B. conceived the project. V.d.B., K.B. and A.P. designed and supervised the project and analyzed the data. V.d.B. performed the genome analyses. A.P. designed and supervised the biochemical experiments with input from V.d.B. K.B. conceived and conducted the structural bioinformatics analysis with input from A.Z. A.P.-T., A.M., J.-L.P., C.B. and T.B. carried out the biochemical experiments. A. Debard, V.P. and M.B.-G. carried out the gene cloning, the protein expression and purifications for the whole collection of MetA and MetX enzymes. A.P. and E.D. designed and analyzed the metabolomics experiments, which were conducted by E.D., P.B. and T.B. C.V.-V. chemically synthesized reference compounds. K.B., A.P. and V.d.B. performed the taxonomic analysis, with input from D.V. and F.A. K.B. built the website with input from V.d.B. V.d.B., K.B. and A.P. wrote the manuscript with input from A. Danchin, M.S., A.Z., C.M., D.V. and J.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–10 and Supplementary Figures 1–22. (PDF 5186 kb)
Supplementary Data Set 1
Primers and strains used for gene cloning. (XLSX 26 kb)
Supplementary Data Set 2
Data from PROCHECK and PROSA II analysis for homology model validation. (XLSX 422 kb)
Rights and permissions
About this article
Cite this article
Bastard, K., Perret, A., Mariage, A. et al. Parallel evolution of non-homologous isofunctional enzymes in methionine biosynthesis. Nat Chem Biol 13, 858–866 (2017). https://doi.org/10.1038/nchembio.2397
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2397
This article is cited by
-
Conversion of methionine biosynthesis in Escherichia coli from trans- to direct-sulfurylation enhances extracellular methionine levels
Microbial Cell Factories (2023)
-
Structural analysis of mycobacterial homoserine transacetylases central to methionine biosynthesis reveals druggable active site
Scientific Reports (2019)
-
Erratum: Corrigendum: Parallel evolution of non-homologous isofunctional enzymes in methionine biosynthesis
Nature Chemical Biology (2017)