Abstract
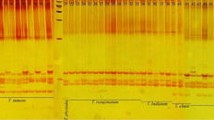
We have identified, isolated, and characterized microsatellite/simple sequence repeat (SSR) loci in trembling aspen (Populus tremuloides) by screening partial genomic libraries. We have also examined the compatibility and use of the P. tremuloides SSR primers to resolve microsatellites in other Populus species. Fourteen microsatellites were identified from 1600 clones screened. The TC/AG microsatellites were the most abundant. A total of 29 alleles were detected in 36 P. tremuloides individuals at the four SSR loci (two each of di- and tri-nucleotide repeats) characterized. The number of alleles at the SSR loci ranged from 5 to 11, with an average of 7.25 alleles per locus, and the observed heterozygosity ranged from 0.19 to 0.82, with a mean of 0.46 per locus. Although the highest polymorphism was observed for a dinucleotide SSR locus, the trinucleotide SSR loci showed substantial polymorphism. There were 34 unique multilocus genotypes among the 36 P. tremuloides individuals examined, and 89% of the individuals had unique multilocus genotypes. Two pairs of SSR primers were successful in PCR, amplifying genomic DNA and resolving microsatellites of comparable size from Populus deltoides, P. nigra, P.×canadensis, and P. maximowiczii. The microsatellite DNA markers developed could be used for clonal fingerprinting, certification of controlled crosses, genome mapping, marker-assisted early selection, genetic diversity assessments, and conservation and sustainable management of poplar genetic resources.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 14 November 1997 / Accepted: 17 November 1997
Rights and permissions
About this article
Cite this article
Dayanandan, S., Rajora, O. & Bawa, K. Isolation and characterization of microsatellites in trembling aspen (Populus tremuloides). Theor Appl Genet 96, 950–956 (1998). https://doi.org/10.1007/s001220050825
Issue Date:
DOI: https://doi.org/10.1007/s001220050825