Abstract
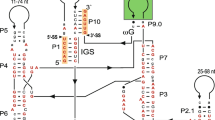
Insertions of less than 100 nt occurring in highly conserved regions of the small subunit ribosomal DNA (SSU rDNA) may represent degenerate forms of the group-I introns observed at the same positions in other organisms. A 63-nt insertion at SSU rDNA position 1512 (relative to theEscherichia coli SSU rDNA) of the lichen-forming fungusArthonia lapidicola can be folded into a secondary structure with two stem loops and a pairing of the insertion and flanking sequences. The two stem loops may correspond to the P1 and P2, and the insertion-flanking pairing to the P10, of a group-I intron. Considering these small insertions as degenerate introns provides important clues to the evolution and catalytic function of group-I introns.
Similar content being viewed by others
References
Armaleo D, Clerc P (1991) Lichen chimeras: DNA analysis suggests that one fungus forms two morphotypes. Exp Mycol 15: 1–10
Burke JM, Esherick JS, Burfeind WR, King JL (1990) A 3′ splice site-binding sequence in the catalytic core of a group-I intron. Nature 344:80–82
Cech TR (1990) Self-splicing of group-I introns. Annu Rev Biochem 59:543–568
Cech TR, Damberger S, Gutell R (1994) Representation of the secondary and tertiary structure of group-I introns. Nature Struct Biol 1:273–280
Christopher DA, Hallick RB (1989)Euglena gracilis chloroplast ribosomal protein operon: a new chloroplast gene for ribosomal protein L5 and description of a novel organelle intron category designated group-III. Nucleic Acids Res 17:7591–7608
Davies RW, Waring RB, Ray JA, Brown TA, Scazzocchio C (1982) Making ends meet: a model for RNA splicing in fungal mitochondria. Nature 300:719–728
DePriest PT (1993) Small subunit rDNA variation in a population of lichen fungi due to optional group-I introns. Gene 134:67–71
Embley TM, Dyal PL, Kilvington S (1992) A group-I intron in the small subunit ribosomal RNA gene fromNaegleria andersoni ssp. andersoni strain PPMFB. Nucleic Acids Res 20:6411
Gargas A, DePriest PT, Grube M, Tehler A (1995 a) Multiple origins of lichen symbioses in fungi suggested by SSU rDNA phylogeny. Science 268:1492–1495
Gargas A, DePriest PT, Taylor JW (1995 b) Positions of multiple insertions in SSU rDNA of lichen-forming fungi. Mol Biol Evol 12:208–218
Garrett RA, Aagaard C, Andersen M, Dalgaard JZ, Lykke-Andersen J, Phan HTN, Trevisanato S, Ostergaard L, Larsen N, Leffers H (1994) Archaeal rRNA operons, intron splicing and horning endonucleases, RNA polymerase operons and phylogeny. System Appl Microbiol 16:680–691
Gast RJ, Fuerst PA, Byers TJ (1994) Discovery of group-I introns in the nuclear small subunit ribosomal RNA genes ofAcanthoamoeba. Nucleic Acids Res 22:592–596
Grube M, DePriest PT, Gargas A, Hafellner J (1995) Isolation of DNA from lichen ascomata. Mycol Res 99:1321–1324
Huss VAR, Hirmer M, Kessler E (1993 a) Genbank accession X73997, X73998
Huss VAR, Wenzeler P, Kessler E (1993 b) Genbank accession X73991
Levra-Juillet E, Boulet A, Seraphin B, Simon M, Faye G (1989) Mitochondrial introns aI1 and/or aI2 are needed for the in vivo deletion of intervening sequences. Mol Gen Genet 217:168–171
Michel F, Westhof E (1990) Modelling of the three-dimensional architecture of group-I catalytic introns based on comparative sequence analysis. J Mol Biol 216:585–610
Michel F, Jacquier JA, Dujon B (1982) Comparison of fungal mitochondrial introns reveals extensive homologies in RNA secondary structure. Biochimie 64:867–881
Michel F, Netter P, Xu M-Q, Shub DA (1990) Mechanism of 3′ splice site selection by the catalytic core of thesun Y intron of bacteriophage T4:the role of a novel base-pairing interaction in group-I introns. Genes Dev 4:777–788
Rogers SO, Yan ZH, Shinohara M, LoBuglio KF, Wang CJK (1993) Messenger RNA intron in the nuclear 18s ribosomal RNA gene of deuteromycetes. Curr Genet 23:338–342
Suh ER, Waring RB (1990) Base pairing between the 3′ exon and an internal guide sequence increases 3′ splice site specificity in theTetrahymena self-splicing rRNA intron. Mol Cell Biol 10:2960–2965
Szostak JW (1986) Enzymatic activity of the conserved core of a group-I self-splicing intron. Nature 322:83–86
Waring RB, Davies RW (1984) Assessment of a model for intron RNA secondary structure relevant to RNA self-splicing — a review. Gene 28:277–291
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In:Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols, a guide to methods and applications. Academic Press, Inc., Harcourt Brace Jovanovich, Publishers, New York, pp 315–322
Wilcox LW, Lewis LA, Fuerst PA, Floyd GL (1991) Group-I introns within the nuclear-encoded small-subunit rRNA gene of three green algae. Mol Biol Evol 9:1103–1118
Author information
Authors and Affiliations
Additional information
Communicated by H. Bertrand
Rights and permissions
About this article
Cite this article
Grube, M., Gargas, A. & DePriest, P.T. A small insertion in the SSU rDNA of the lichen fungusArthonia lapidicola is a degenerate group-I intron. Curr Genet 29, 582–586 (1996). https://doi.org/10.1007/BF02426963
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF02426963