Abstract
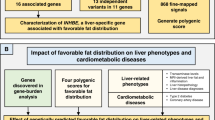
Body-fat distribution is a risk factor for adverse cardiovascular health consequences. We analyzed the association of body-fat distribution, assessed by waist-to-hip ratio adjusted for body mass index, with 228,985 predicted coding and splice site variants available on exome arrays in up to 344,369 individuals from five major ancestries (discovery) and 132,177 European-ancestry individuals (validation). We identified 15 common (minor allele frequency, MAF ≥5%) and nine low-frequency or rare (MAF <5%) coding novel variants. Pathway/gene set enrichment analyses identified lipid particle, adiponectin, abnormal white adipose tissue physiology and bone development and morphology as important contributors to fat distribution, while cross-trait associations highlight cardiometabolic traits. In functional follow-up analyses, specifically in Drosophila RNAi-knockdowns, we observed a significant increase in the total body triglyceride levels for two genes (DNAH10 and PLXND1). We implicate novel genes in fat distribution, stressing the importance of interrogating low-frequency and protein-coding variants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Summary statistics of all analyses are available at https://portals.broadinstitute.org/collaboration/giant/index.php/GIANT_consortium_data_files.
Change history
07 March 2019
In the HTML version of the article originally published, the link for Supplementary Data 5 returned the file for Supplementary Data 7. The error has been corrected in the HTML version of the article.
References
Pischon, T. et al. General and abdominal adiposity and risk of death in Europe. N. Engl. J. Med. 359, 2105–2120 (2008).
Wang, Y., Rimm, E. B., Stampfer, M. J., Willett, W. C. & Hu, F. B. Comparison of abdominal adiposity and overall obesity in predicting risk of type 2 diabetes among men. Am. J. Clin. Nutr. 81, 555–563 (2005).
Canoy, D. Distribution of body fat and risk of coronary heart disease in men and women. Curr. Opin. Cardiol. 23, 591–598 (2008).
Snijder, M. B. et al. Associations of hip and thigh circumferences independent of waist circumference with the incidence of type 2 diabetes: the Hoorn Study. Am. J. Clin. Nutr. 77, 1192–1197 (2003).
Yusuf, S. et al. Obesity and the risk of myocardial infarction in 27,000 participants from 52 countries: a case-control study. Lancet 366, 1640–1649 (2005).
Mason, C., Craig, C. L. & Katzmarzyk, P. T. Influence of central and extremity circumferences on all-cause mortality in men and women. Obesity. 16, 2690–2695 (2008).
Karpe, F. & Pinnick, K. E. Biology of upper-body and lower-body adipose tissue—llink to whole-body phenotypes. Nat. Rev. Endocrinol. 11, 90–100 (2015).
Manolopoulos, K. N., Karpe, F. & Frayn, K. N. Gluteofemoral body fat as a determinant of metabolic health. Int. J. Obes. 34, 949–959 (2010).
Emdin, C. A. et al. Genetic association of waist-to-hip ratio with cardiometabolic traits, type 2 diabetes, and coronary heart disease. JAMA 317, 626–634 (2017).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Winkler, T. W. et al. The influence of age and sex on genetic associations with adult body size and shape: a large-scale genome-wide interaction study. PLoS Genet. 11, e1005378 (2015).
Wen, W. et al. Genome-wide association studies in east asians identify new loci for waist-hip ratio and waist circumference. Sci. Rep. 6, 17958 (2016).
Gao, C. et al. A comprehensive analysis of common and rare variants to identify adiposity loci in hispanic Americans: the iras family study (IRASFS). PLoS ONE 10, e0134649 (2015).
Graff, M. et al. Genome-wide physical activity interactions in adiposity—meta-analysis of 200,452 adults. PLoS Genet. 13, e1006528 (2017).
Justice, A. E. et al. Genome-wide meta-analysis of 241,258 adults accounting for smoking behaviour identifies novel loci for obesity traits. Nat. Commun. 8, 14977 (2017).
Ng, M. C. Y. et al. Discovery and fine-mapping of adiposity loci using high density imputation of genome-wide association studies in individuals of African ancestry: African Ancestry Anthropometry Genetics Consortium. PLoS Genet. 13, e1006719 (2017).
Aschard, H., Vilhjalmsson, B. J., Joshi, A. D., Price, A. L. & Kraft, P. Adjusting for heritable covariates can bias effect estimates in genome-wide association studies. Am. J. Hum. Genet. 96, 329–339 (2015).
Day, F. R., Loh, P. R., Scott, R. A., Ong, K. K. & Perry, J. R. A robust example of collider bias in a genetic association study. Am. J. Hum. Genet. 98, 392–393 (2016).
Feng, S., Liu, D., Zhan, X., Wing, M. K. & Abecasis, G. R. RAREMETAL: fast and powerful meta-analysis for rare variants. Bioinformatics 30, 2828–2829 (2014).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Marouli, E. et al. Rare and low-frequency coding variants alter human adult height. Nature 542, 186–190 (2017).
Lamparter, D., Marbach, D., Rueedi, R., Kutalik, Z. & Bergmann, S. Fast and rigorous computation of gene and pathway scores from SNP-based summary statistics. PLoS Comput. Biol. 12, e1004714 (2016).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Kawai, M., de Paula, F. J. & Rosen, C. J. New insights into osteoporosis: the bone-fat connection. J. Intern. Med. 272, 317–329 (2012).
Turcot, V. et al. Protein-altering variants associated with body mass index implicate pathways that control energy intake and expenditure in obesity. Nat. Genet. 50, 26–41 (2018).
Liu, D. J. et al. Exome-wide association study of plasma lipids in >300,000 individuals. Nat. Genet. 49, 1758–1766 (2017).
Kraja, A. T. et al. New blood pressure-associated loci identified in meta-analyses of 475 000 individuals. Circ. Cardiovasc. Genet. 10, e001778 (2017).
Mahajan, A. et al. Identification and functional characterization of G6PC2 coding variants influencing glycemic traits define an effector transcript at the G6PC2-ABCB11 locus. PLoS Genet. 11, e1004876 (2015).
Manning, A. et al. A low-frequency inactivating akt2 variant enriched in the finnish population is associated with fasting insulin levels and type 2 diabetes risk. Diabetes 66, 2019–2032 (2017).
Zhao, W. et al. Identification of new susceptibility loci for type 2 diabetes and shared etiological pathways with coronary heart disease. Nat. Genet. 49, 1450–1457 (2017).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Ng, M. C. et al. Meta-analysis of genome-wide association studies in African Americans provides insights into the genetic architecture of type 2 diabetes. PLoS Genet. 10, e1004517 (2014).
Mahajan, A. et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat. Genet. 46, 234–244 (2014).
Saxena, R. et al. Genome-wide association study identifies a novel locus contributing to type 2 diabetes susceptibility in Sikhs of Punjabi origin from India. Diabetes 62, 1746–1755 (2013).
Cook, J. P. & Morris, A. P. Multi-ethnic genome-wide association study identifies novel locus for type 2 diabetes susceptibility. Eur. J. Hum. Genet. 24, 1175–1180 (2016).
Voight, B. F. et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat. Genet. 42, 579–589 (2010).
Burdett, T. et al. The NHGRI-EBI Catalog of published genome-wide association studies. V.1.0 edn Vol. 2015 (NHGRI-EBI, 2015).
Hindorff, L. A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl Acad. Sci. USA 106, 9362–9367 (2009).
Lutoslawska, G. et al. Relationship between the percentage of body fat and surrogate indices of fatness in male and female Polish active and sedentary students. J. Physiol. Anthropol. 33, 10 (2014).
Verma, M., Rajput, M., Sahoo, S. S., Kaur, N. & Rohilla, R. Correlation between the percentage of body fat and surrogate indices of obesity among adult population in rural block of Haryana. J. Family Med. Prim. Care 5, 154–159 (2016).
Pereira, P. F. et al. Measurements of location of body fat distribution: an assessment of colinearity with body mass, adiposity and stature in female adolescents. Rev. Paul. Pediatr. 33, 63–71 (2015).
Lu, Y. et al. New loci for body fat percentage reveal link between adiposity and cardiometabolic disease risk. Nat. Commun. 7, 10495 (2016).
Chambers, J. C. et al. Common genetic variation near MC4R is associated with waist circumference and insulin resistance. Nat. Genet. 40, 716–718 (2008).
Nead, K. T. et al. Contribution of common non-synonymous variants in PCSK1 to body mass index variation and risk of obesity: a systematic review and meta-analysis with evidence from up to 331 175 individuals. Hum. Mol. Genet. 24, 3582–3594 (2015).
Pospisilik, J. A. et al. Drosophila genome-wide obesity screen reveals hedgehog as a determinant of brown versus white adipose cell fate. Cell 140, 148–160 (2010).
Consortium, G. T. Human genomics. The genotype-tissue expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Baraille, F., Planchais, J., Dentin, R., Guilmeau, S. & Postic, C. Integration of chrebp-mediated glucose sensing into whole body metabolism. Physiology 30, 428–437 (2015).
Kursawe, R. et al. Decreased transcription of ChREBP-alpha/beta isoforms in abdominal subcutaneous adipose tissue of obese adolescents with prediabetes or early type 2 diabetes: associations with insulin resistance and hyperglycemia. Diabetes 62, 837–844 (2013).
Lotta, L. A. et al. Integrative genomic analysis implicates limited peripheral adipose storage capacity in the pathogenesis of human insulin resistance. Nat. Genet. 49, 17–26 (2017).
Cargill, M. et al. A large-scale genetic association study confirms IL12B and leads to the identification of IL23R as psoriasis-risk genes. Am. J. Hum. Genet. 80, 273–290 (2007).
Hazlett, J. et al. IL-23R rs11209026 polymorphism modulates IL-17A expression in patients with rheumatoid arthritis. Genes Immun. 13, 282–287 (2012).
Karaderi, T. et al. Association between the interleukin 23 receptor and ankylosing spondylitis is confirmed by a new UK case-control study and meta-analysis of published series. Rheumatology 48, 386–389 (2009).
Duerr, R. H. et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 314, 1461–1463 (2006).
Abdollahi, E., Tavasolian, F., Momtazi-Borojeni, A. A., Samadi, M. & Rafatpanah, H. Protective role of R381Q (rs11209026) polymorphism in IL-23R gene in immune-mediated diseases: comprehensive review. J. Immunotoxicol. 13, 286–300 (2016).
Abraham, C., Dulai, P. S., Vermeire, S. & Sandborn, W. J. Lessons learned from trials targeting cytokine pathways in patients with inflammatory bowel diseases. Gastroenterology 152, 374–388 e4 (2017).
Molinelli, E., Campanati, A., Ganzetti, G. & Offidani, A. Biologic therapy in immune mediated inflammatory disease: basic science and clinical concepts. Curr. Drug Saf. 11, 35–43 (2016).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47 (2016).
Wells, J. C. Sexual dimorphism of body composition. Best. Pract. Res. Clin. Endocrinol. Metab. 21, 415–430 (2007).
Loomba-Albrecht, L. A. & Styne, D. M. Effect of puberty on body composition. Curr. Opin. Endocrinol. Diabetes. Obes. 16, 10–15 (2009).
Rogol, A. D., Roemmich, J. N. & Clark, P. A. Growth at puberty. J. Adolesc. Health 31, 192–200 (2002).
Gibson, G. Rare and common variants: twenty arguments. Nat. Rev. Genet. 13, 135–145 (2012).
Stern, J. H., Rutkowski, J. M. & Scherer, P. E. Adiponectin, leptin, and fatty acids in the maintenance of metabolic homeostasis through adipose tissue crosstalk. Cell. Metab. 23, 770–784 (2016).
Dewey, F. E. et al. Inactivating variants in angptl4 and risk of coronary artery disease. N. Engl. J. Med. 374, 1123–1133 (2016).
Bondestam, J. et al. cDNA cloning, expression studies and chromosome mapping of human type I serine/threonine kinase receptor ALK7 (ACVR1C). Cytogenet. Cell Genet. 95, 157–162 (2001).
Jornvall, H., Blokzijl, A., ten Dijke, P. & Ibanez, C. F. The orphan receptor serine/threonine kinase ALK7 signals arrest of proliferation and morphological differentiation in a neuronal cell line. J. Biol. Chem. 276, 5140–5146 (2001).
Kim, B. C. et al. Activin receptor-like kinase-7 induces apoptosis through activation of MAPKs in a Smad3-dependent mechanism in hepatoma cells. J. Biol. Chem. 279, 28458–28465 (2004).
Watanabe, R. et al. The MH1 domains of smad2 and smad3 are involved in the regulation of the ALK7 signals. Biochem. Biophys. Res. Commun. 254, 707–712 (1999).
Kogame, M. et al. ALK7 is a novel marker for adipocyte differentiation. J. Med. Invest. 53, 238–245 (2006).
Murakami, M. et al. Expression of activin receptor-like kinase 7 in adipose tissues. Biochem. Genet. 51, 202–210 (2013).
Carlsson, L. M. et al. ALK7 expression is specific for adipose tissue, reduced in obesity and correlates to factors implicated in metabolic disease. Biochem. Biophys. Res. Commun. 382, 309–314 (2009).
Carithers, L. J. & Moore, H. M. The genotype-tissue expression (GTEx) project. Biopreserv. Biobank. 13, 307–308 (2015).
Yogosawa, S., Mizutani, S., Ogawa, Y. & Izumi, T. Activin receptor-like kinase 7 suppresses lipolysis to accumulate fat in obesity through downregulation of peroxisome proliferator-activated receptor gamma and C/EBPalpha. Diabetes 62, 115–123 (2013).
Yogosawa, S. & Izumi, T. Roles of activin receptor-like kinase 7 signaling and its target, peroxisome proliferator-activated receptor gamma, in lean and obese adipocytes. Adipocyte 2, 246–250 (2013).
Seifi, M., Ghasemi, A., Namipashaki, A. & Samadikuchaksaraei, A. Is C771G polymorphism of MLX interacting protein-like (MLXIPL) gene a novel genetic risk factor for non-alcoholic fatty liver disease? Cell Mol. Biol. 60, 37–42 (2014).
Cairo, S., Merla, G., Urbinati, F., Ballabio, A. & Reymond, A. WBSCR14, a gene mapping to the Williams–Beuren syndrome deleted region, is a new member of the Mlx transcription factor network. Hum. Mol. Genet. 10, 617–627 (2001).
Ambele, M. A., Dessels, C., Durandt, C. & Pepper, M. S. Genome-wide analysis of gene expression during adipogenesis in human adipose-derived stromal cells reveals novel patterns of gene expression during adipocyte differentiation. Stem Cell Res. 16, 725–734 (2016).
Marchler-Bauer, A. et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 43, D222–D226 (2015).
Toyofuku, T. et al. Semaphorin-4A, an activator for T-cell-mediated immunity, suppresses angiogenesis via Plexin-D1. EMBO J. 26, 1373–1384 (2007).
Gitler, A. D., Lu, M. M. & Epstein, J. A. PlexinD1 and semaphorin signaling are required in endothelial cells for cardiovascular development. Dev. Cell. 7, 107–116 (2004).
Luchino, J. et al. Semaphorin 3E suppresses tumor cell death triggered by the plexin D1 dependence receptor in metastatic breast cancers. Cancer Cell. 24, 673–685 (2013).
Shimizu, I. et al. Semaphorin3E-induced inflammation contributes to insulin resistance in dietary obesity. Cell. Metab. 18, 491–504 (2013).
Verzijl, H. T., van der Zwaag, B., Cruysberg, J. R. & Padberg, G. W. Mobius syndrome redefined: a syndrome of rhombencephalic maldevelopment. Neurology 61, 327–333 (2003).
Verzijl, H. T., van der Zwaag, B., Lammens, M., ten Donkelaar, H. J. & Padberg, G. W. The neuropathology of hereditary congenital facial palsy vs Mobius syndrome. Neurology 64, 649–653 (2005).
Fujita, M., Reinhart, F. & Neutra, M. Convergence of apical and basolateral endocytic pathways at apical late endosomes in absorptive cells of suckling rat ileum in vivo. J. Cell Sci. 97(Pt 2), 385–394 (1990).
Briegel, W. Neuropsychiatric findings of mobius sequence—a review. Clin. Genet. 70, 91–97 (2006).
Ta-Shma, A. et al. Isolated truncus arteriosus associated with a mutation in the plexin-D1 gene. Am. J. Med. Genet. A. 161A, 3115–3120 (2013).
Mazzotta, C. et al. Plexin-D1/Semaphorin 3E pathway may contribute to dysregulation of vascular tone control and defective angiogenesis in systemic sclerosis. Arthritis Res. Ther. 17, 221 (2015).
Yang, W. J. et al. Semaphorin-3C signals through Neuropilin-1 and PlexinD1 receptors to inhibit pathological angiogenesis. EMBO Mol. Med. 7, 1267–1284 (2015).
Zygmunt, T. et al. Semaphorin-PlexinD1 signaling limits angiogenic potential via the VEGF decoy receptor sFlt1. Dev. Cell. 21, 301–314 (2011).
Kim, J., Oh, W. J., Gaiano, N., Yoshida, Y. & Gu, C. Semaphorin 3E-Plexin-D1 signaling regulates VEGF function in developmental angiogenesis via a feedback mechanism. Genes Dev. 25, 1399–1411 (2011).
Bertolino, P. et al. Activin B receptor ALK7 is a negative regulator of pancreatic beta-cell function. Proc. Natl Acad. Sci. USA 105, 7246–7251 (2008).
Haworth, K. E. et al. Methylation of the FGFR2 gene is associated with high birth weight centile in humans. Epigenomics 6, 477–491 (2014).
Chi, X. et al. Angiopoietin-like 4 modifies the interactions between lipoprotein lipase and its endothelial cell transporter GPIHBP1. J. Biol. Chem. 290, 11865–11877 (2015).
Catoire, M. et al. Fatty acid-inducible ANGPTL4 governs lipid metabolic response to exercise. Proc. Natl Acad. Sci. USA 111, E1043–E1052 (2014).
van Raalte, D. H. et al. Angiopoietin-like protein 4 is differentially regulated by glucocorticoids and insulin in vitro and in vivo in healthy humans. Exp. Clin. Endocrinol. Diabetes. 120, 598–603 (2012).
Koster, A. et al. Transgenic angiopoietin-like (angptl)4 overexpression and targeted disruption of angptl4 and angptl3: regulation of triglyceride metabolism. Endocrinology 146, 4943–4950 (2005).
Thiagalingam, A. et al. RREB-1, a novel zinc finger protein, is involved in the differentiation response to Ras in human medullary thyroid carcinomas. Mol. Cell. Biol. 16, 5335–5345 (1996).
Bonomo, J. A. et al. The ras responsive transcription factor RREB1 is a novel candidate gene for type 2 diabetes associated end-stage kidney disease. Hum. Mol. Genet. 23, 6441–6447 (2014).
Thiagalingam, A., Lengauer, C., Baylin, S. B. & Nelkin, B. D. RREB1, a ras responsive element binding protein, maps to human chromosome 6p25. Genomics 45, 630–632 (1997).
Bisogno, T. et al. Cloning of the first sn1-DAG lipases points to the spatial and temporal regulation of endocannabinoid signaling in the brain. J. Cell Biol. 163, 463–468 (2003).
Global Lipids Genetics, C. et al. Discovery and refinement of loci associated with lipid levels. Nat. Genet. 45, 1274–1283 (2013).
Kooner, J. S. et al. Genome-wide scan identifies variation in MLXIPL associated with plasma triglycerides. Nat. Genet. 40, 149–151 (2008).
Pan, L. A. et al. G771C polymorphism in the mlxipl gene is associated with a risk of coronary artery disease in the chinese: a case-control study. Cardiology 114, 174–178 (2009).
Kang, G., Leech, C. A., Chepurny, O. G., Coetzee, W. A. & Holz, G. G. Role of the cAMP sensor Epac as a determinant of KATP channel ATP sensitivity in human pancreatic beta-cells and rat INS-1 cells. J. Physiol. 586, 1307–1319 (2008).
Ji, Z., Mei, F. C. & Cheng, X. Epac, not PKA catalytic subunit, is required for 3T3-L1 preadipocyte differentiation. Front Biosci. 2, 392–398 (2010).
Martini, C. N., Plaza, M. V. & Vila Mdel, C. PKA-dependent and independent cAMP signaling in 3T3-L1 fibroblasts differentiation. Mol. Cell. Endocrinol. 298, 42–47 (2009).
Petersen, R. K. et al. Cyclic AMP (cAMP)-mediated stimulation of adipocyte differentiation requires the synergistic action of Epac- and cAMP-dependent protein kinase-dependent processes. Mol. Cell. Biol. 28, 3804–3816 (2008).
Yan, J. et al. Enhanced leptin sensitivity, reduced adiposity, and improved glucose homeostasis in mice lacking exchange protein directly activated by cyclic AMP isoform 1. Mol. Cell. Biol. 33, 918–926 (2013).
Gesta, S. et al. Evidence for a role of developmental genes in the origin of obesity and body fat distribution. Proc. Natl Acad. Sci. USA 103, 6676–6681 (2006).
Gesta, S. et al. Mesodermal developmental gene Tbx15 impairs adipocyte differentiation and mitochondrial respiration. Proc. Natl Acad. Sci. USA 108, 2771–2776 (2011).
Lee, K. Y. et al. Tbx15 controls skeletal muscle fibre-type determination and muscle metabolism. Nat. Commun. 6, 8054 (2015).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Liu, D. J. et al. Meta-analysis of gene-level tests for rare variant association. Nat. Genet. 46, 200–204 (2014).
Goldstein, J. I. et al. zCall: a rare variant caller for array-based genotyping: genetics and population analysis. Bioinformatics 28, 2543–2545 (2012).
Winkler, T. W. et al. Quality control and conduct of genome-wide association meta-analyses. Nat. Protoc. 9, 1192–1212 (2014).
Winkler, T. W. et al. EasyStrata: evaluation and visualization of stratified genome-wide association meta-analysis data. Bioinformatics 31, 259–261 (2015).
Purcell, S. M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Yang, J. et al. Genomic inflation factors under polygenic inheritance. Eur. J. Hum. Genet. 19, 807–812 (2011).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, S1–S3 (2012).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Wellcome Trust Case Control, C. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Frey, B. J. & Dueck, D. Clustering by passing messages between data points. Science 315, 972–976 (2007).
Moayyeri, A., Hammond, C. J., Valdes, A. M. & Spector, T. D. Cohort profile: TwinsUK and healthy ageing twin study. Int. J. Epidemiol. 42, 76–85 (2013).
Boyd, A. et al. Cohort profile: the ‘children of the 90s’–the index offspring of the Avon Longitudinal Study of parents and children. Int. J. Epidemiol. 42, 111–127 (2013).
Kutalik, Z., Whittaker, J., Waterworth, D., Beckmann, J. S. & Bergmann, S. Novel method to estimate the phenotypic variation explained by genome-wide association studies reveals large fraction of the missing heritability. Genet. Epidemiol. 35, 341–349 (2011).
Billingsley, P. Probability and Measure (Wiley, New York, 1986).
Surendran, P. et al. Trans-ancestry meta-analyses identify rare and common variants associated with blood pressure and hypertension. Nat. Genet. 48, 1151–1161 (2016).
Nikpay, M. et al. A comprehensive 1000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet. 47, 1121–1130 (2015).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003).
Civelek, M. et al. Genetic regulation of adipose gene expression and cardio-metabolic traits. Am. J. Hum. Genet. 100, 428–443 (2017).
Acknowledgements
This work was primarily supported through funding from the National Institute of Health (NIH): 1K99HL130580, R01-DK089256, 2R01HD057194, U01HG007416, R01DK101855, T32 HL007055, KL2TR001109; and the American Heart Association (AHA): 13POST16500011 and 13GRNT16490017. Co-author Y. Jia recently passed away while this work was in process. This study was completed as part of the Genetic Investigation of ANtropometric Traits (GIANT) Consortium. This research has been conducted using the UK Biobank resource. A full list of acknowledgements is provided in the Supplementary Data 18.
Author information
Authors and Affiliations
Consortia
Contributions
Writing Group: L.A.C., R.S.F., T.M.F., M. Graff, H.M.H., J.N.H., A.E.J., T.K., Z.K., C.M.L., R.J.F.L., Y.L., K.E.N., V.T., K.L.Y.
Data preparation group: T.A., I.B.B., T.E., S.F., M. Graff, H.M.H., A.E.J., T.K., D.J.L., K.S.L., A.E.L., R.J.F.L., Y.L., E. Marouli, N.G.D.M., C.M.-G, P.M., M.C.Y.N., M.A.R., S.S., C.S., K. Stirrups, V.T., S.V., S.M.W., T.W.W., K.L.Y., X.Z.
WHR meta-analyses: P.L.A., H.M.H., A.E.J., T.K., M. Graff, C.M.L., R.J.F.L., K.E.N., V.T., K.L.Y.
Pleiotropy working group: G.A., M. Boehnke, J.P.C., P.D., F.D., J.C.F., H.M.H., H.K., H.M.H., A.E.J., C.M.L., D.J.L., R.J.F.L., A. Mahajan, E. Marouli, G.M., M.I.M., P.B.M., G.M.P., J.R.B.P., K.S.R., X.S., S.W., J.W., C.J.W.
Phenome-wide association studies: L. Bastarache, J.C.D., A.G., A. Mahajan, M.I.M.
Gene-set enrichment analyses: S.B., R.S.F., J.N.H., Z.K., D.L., T.H.P., T.F.V.
eQTL analyses: C.K.R., Y.L., K.L.M.
Monogenic and syndromic gene enrichment analyses: H.M.H., A.K.M.
Fly Obesity Screen: A. Lempradl, J.A. Pospisilik.
Overseeing of contributing studies and consortia: (1958 Birth Cohort) P.D.; (Airwave) P.E.; (AMC PAS) G.K.H.; (Amish) J.R.O.; (ARIC) E.B.; (ARIC, Add Health) K.E.N.; (BRAVE) E.D.A., R.C.; (BRIGHT) P.B.M.; (CARDIA) M.F., P.J.S.; (Cebu Longitudinal Health and Nutrition Survey) K.L.M.; (CHD Exome + Consortium) A.S.B., J.M.M.H., D.F.R., J.D.; (CHES) R.V.; (Clear/eMERGE (Seattle)) G.P.J.; (CROATIA_Korcula) V.V., O. Polasek, I.R.; (deCODE) K. Stefansson, U.T.; (DHS) D.W.B.; (DIACORE) C.A.B.; (DPS) J.T., J. Lindström, M.U.; (DRSEXTRA) T.A.L., R.R.; (EFSOCH) A.T.H., T.M.F.; (EGCUT) T.E.; (eMERGE (Seattle)) E.B.L.; (EPIC-Potsdam) M.B.S., H.B.; (EpiHealth) E.I., P.W.F.; (EXTEND) A.T.H., T.M.F.; (Family Heart Study) I.B.B.; (Fenland, EPIC) R.A.S.; (Fenland, EPIC, InterAct) N.J.W., C.L.; (FINRISK) S. Männistö; (FINRISK 2007 (T2D)) P.J., V. Salomaa; (Framingham Heart Study) L.A.C.; (FUSION) M. Boehnke, F.S.C.; (FVG) P.G.; (Generation Scotland) C.H., B.H.S.; (Genetic Epidemiology Network of Arteriopathy (GENOA)) S.L.R.K.; (GRAPHIC) N.J.S.; (GSK-STABILITY) D.M.W., L.W., H.D.W.; (Health) A. Linneberg; (HELIC MANOLIS) E.Z., G.D.; (HELIC Pomak) E.Z., G.D.; (HUNT-MI) K.H., C.J.W.; (Inter99) T.H., T.J.; (IRASFS) L.E.W., E.K.S.; (Jackson Heart Study (JHS)) J.G.W.; (KORA S4) K. Strauch, I.M.H.; (Leipzig-Adults) M. Blüher, P. Kovacs.; (LOLIPOP-Exome) J.C.C., J.S.K.; (LOLIPOP-OmniEE) J.C.C., J.S.K.; (MESA) J.I.R., X.G.; (METSIM) J.K., M.L.; (MONICA-Brianza) G.C.; (Montreal Heart Institute Biobank (MHIBB)) M.P.D., G.L., S.d.D., J.C.T.; (MORGAM Central Laboratory) M.P.; (MORGAM Data Centre) K.K.; (OBB) F. Karpe; (PCOS) A.P.M., C.M.L.; (PIVUS) C.M.L., L.L.; (PRIME—Belfast) F. Kee; (PRIME—Lille) P.A.; (PRIME—Strasbourg) M.M.; (PRIME—Toulouse) J.F.; (PROMIS) D.S.; (quality control) M.A.R.; (RISC) B.B., E.F., M.W.; (Rotterdam Study I) A.G.U., M.A.I.; (SEARCH) A.M.D.; (SHIP/SHIP-Trend) M.D.; (SIBS) D.F.E.; (SOLID TIMI-52) D.M.W.; (SORBS) A.P.M., M.S., A. Tönjes; (The Mount Sinai BioMe Biobank) E.P.B., R.J.F.L.; (The NEO Study) D.O.M.K.; (The NHAPC Study, The GBTDS Study) X.L.; (The Western Australian Pregnancy Cohort (Raine) Study) C.E.P., S. Macgregor; (TwinsUK) T.D.S.; (ULSAM) A.P.M.; (Vejle Biobank) I.B., C.C., O. Pedersen; (WGHS) D.I.C., P.M.R.; (Women’s Health Initiative) P.L.A.; (WTCCC-UKT2D) M.I.M., K.R.O.; (YFS) T.L., O.T.R.
Genotyping of contributing studies and consortia: (1958 Birth Cohort) K.E.S.; (Airwave) E.E., M.S.L.; (AMC PAS) S.S.; (Amish) L.M.Y.A., J.A. Perry; (ARIC) E.W.D., M.L.G.; (BBMRI-NL) S.H.V., L. Broer, C.M.v.D., P.I.W.d.B.; (BRAVE) E.D.A.; (Cambridge Cancer Studies) J.G.D.; (CARDIA) M.F.; (CHD Exome + Consortium) A.S.B., J.M.M.H., D.F.R., J.D., R.Y.; (Clear/eMERGE (Seattle)) G.P.J.; (CROATIA_Korcula) V.V.; (DIACORE) C.A.B., M. Gorski; (DPS) A.U.J., J. Lindström; (DRSEXTRA) P. Komulainen; (EGCUT) T.E.; (EPIC-Potsdam) M.B.S., K.M.; (EpiHealth) E.I., P.W.F.; (Family Heart Study) K.D.T.; (Fenland, EPIC) R.A.S.; (Fenland, EPIC, InterAct) N.J.W., C.L.; (FUSION) N.N.; (FVG) I.G., A. Morgan; (Generation Scotland) C.H.; (Genetic Epidemiology Network of Arteriopathy (GENOA)) S.L.R.K., J.A.S.; (GRAPHIC) N.J.S.; (GSK-STABILITY) D.M.W.; (Health) J.B.J.; (HELIC MANOLIS) L. Southam; (HELIC Pomak) L. Southam; (Inter99) T.H., N.G.; (KORA) M.M.N.; (KORA S4) K. Strauch, H.G.; (Leipzig-Adults) A. Mahajan; (LOLIPOP-Exome) J.C.C., J.S.K.; (LOLIPOP-OmniEE) J.C.C., J.S.K.; (MESA) J.I.R., Y.D.I.C., K.D.T.; (METSIM) J.K., M.L.; (Montreal Heart Institute Biobank (MHIBB)) M.P.D.; (OBB) F. Karpe; (PCOS) A.P.M.; (PIVUS) C.M.L.; (Rotterdam Study I) A.G.U., C.M.G., F.R.; (SDC) J.M.J., H.V.; (SEARCH) A.M.D.; (SOLID TIMI-52) D.M.W.; (SORBS) A.P.M.; (The Mount Sinai BioMe Biobank) E.P.B., R.J.F.L., Y.L., C.S.; (The NEO Study) R.L.G.; (The NHAPC study, The GBTDS study) X.L., H.L., Y.H.; (The Western Australian Pregnancy Cohort (Raine) Study) C.E.P., S. Macgregor; (TUDR) Z.A.; (TwinsUK) A.P.M.; (ULSAM) A.P.M.; (WGHS) D.I.C., A.Y.C.; (Women’s Health Initiative) A.P.R.; (WTCCC-UKT2D) M.I.M.; (YFS) T.L., L.P.L.
Phenotyping of contributing studies and consortia: (Airwave) E.E.; (AMC PAS) S.S.; (Amish) L.M.Y.A.; (ARIC) E.W.D.; (ARIC, Add Health) K.E.N.; (BBMRI-NL) S.H.V.; (BRAVE) E.D.A.; (BRIGHT) M.J.C.; (CARL) A. Robino, G.G.; (Cebu Longitudinal Health and Nutrition Survey) N.R.L.; (CHES) R.V., M.T.; (Clear/eMERGE (Seattle)) G.P.J., A.A.B.; (CROATIA_Korcula) O. Polasek, I.R.; (DIACORE) C.A.B., B.K.K.; (DPS) A.U.J., J. Lindström; (EFSOCH) A.T.H.; (EGCUT) E. Mihailov; (EPIC-Potsdam) H.B.; (EpiHealth) E.I.; (EXTEND) A.T.H.; (Family Heart Study) M.F.F.; (Fenland, EPIC, InterAct) N.J.W.; (FIN-D2D 2007) L.M., M.V.; (FINRISK) S. Männistö; (FINRISK 2007 (T2D)) P.J., H.M.S.; (Framingham Heart Study) C.S.F.; (Generation Scotland) C.H., B.H.S.; (Genetic Epidemiology Network of Arteriopathy (GENOA)) S.L.R.K., J.A.S.; (GRAPHIC) N.J.S.; (GSK-STABILITY) L.W., H.D.W.; (Health) A. Linneberg, B.H.T.; (HELIC MANOLIS) L. Southam, A.E.F., E.T.; (HELIC Pomak) L. Southam, A.E.F., M.K.; (HUNT-MI) K.H., O.L.H.; (Inter99) T.J., N.G.; (IRASFS) L.E.W., B.K.; (KORA) M.M.N.; (BBMRI-NL) K.M.A.S.; (Leipzig-Adults) M. Blüher, P. Kovacs; (LOLIPOP-Exome) J.C.C., J.S.K.; (LOLIPOP-OmniEE) J.C.C., J.S.K.; (MESA) M.A.; (Montreal Heart Institute Biobank (MHIBB)) G.L., K.S.L., V.T.; (MORGAM Data Centre) K.K.; (OBB) F. Karpe, M.N.; (PCOS) C.M.L.; (PIVUS) L.L.; (PRIME—Belfast) F. Kee; (PRIME—Lille) P.A.; (PRIME—Strasbourg) M.M.; (PRIME—Toulouse) J.F.; (RISC) B.B., E.F.; (Rotterdam Study I) M.A.I., C.M.G., F.R., M.C.Z.; (SHIP/SHIP-Trend) N.F.; (SORBS) M.S., A. Tönjes; (The Mount Sinai BioMe Biobank) E.P.B., Y.L., C.S.; (The NEO Study) R.d.M.; (The NHAPC study, The GBTDS study) X.L., H.L., L. Sun, F.W.; (The Western Australian Pregnancy Cohort (Raine) Study) C.E.P.; (TUDR) Y.J.H., W.J.L.; (TwinsUK) T.D.S., K.S.S.; (ULSAM) V.G.; (WGHS) D.I.C., P.M.R.; (Women’s Health Initiative) A.P.R.; (WTCCC-UKT2D) M.I.M., K.R.O.; (YFS) T.L., O.T.R.
Data analysis of contributing studies and consortia: (1958 Birth Cohort) K.E.S., I.N.; (Airwave) E.E., M.S.L.; (AMC PAS) S.S.; (Amish) J.R.O., L.M.Y.A., J.A. Perry; (ARIC, Add Health) K.E.N., K.L.Y., M. Graff; (BBMRI-NL) L. Broer; (BRAVE) R.C., D.S.A.; (BRIGHT) H.R.W.; (Cambridge Cancer Studies) J.G.D., A.E., D.J.T.; (CARDIA) C.E.L, M.F., L.-A.L.; (CARL) A. Robino, D.V.; (Cebu Longitudinal Health and Nutrition Survey) Y.W.; (CHD Exome + Consortium) A.S.B., J.M.M.H., D.F.R., R.Y., P.S.; (CHES) Y.J.; (CROATIA_Korcula) V.V.; (deCODE) V. Steinthorsdottir, G.T.; (DHS) A.J.C., P.M., M.C.Y.N.; (DIACORE) C.A.B., M. Gorski; (EFSOCH) H.Y.; (EGCUT) T.E., R.M.; (eMERGE (Seattle)) D.S.C.; (ENDO) T.K.; (EPIC) J.H.Z.; (EPIC-Potsdam) K.M.; (EpiHealth) S.G.; (EXTEND) H.Y.; (Family Heart Study) M.F.F.; (Fenland) J.Luan.; (Fenland, EPIC) R.A.S.; (Fenland, InterAct) S.M.W.; (FIN-D2D 2007) M.V., L.M.; (Finrisk Extremes and quality control) S.V.; (Framingham Heart Study) C.T.L., N.L.H.C.; (FVG) I.G.; (Generation Scotland) C.H., J.M.; (Genetic Epidemiology Network of Arteriopathy (GENOA)) L.F.B.; (GIANT-Analyst) A.E.J.; (GRAPHIC) N.J.S., N.G.D.M., C.P.N.; (GSK-STABILITY) D.M.W., A.J.S.; (Health) J.B.J.; (HELIC MANOLIS) L. Southam; (HELIC Pomak) L. Southam; (HUNT-MI) W. Zhou; (Inter99) N.G.; (IRASFS) B.K.; (Jackson Heart Study (JHS)) L.A.L., J. Li; (KORA S4) T.W.W.; (BBMRI-NL) K.M.A.S.; (Leipzig-Adults) A. Mahajan; (LOLIPOP-Exome) J.C.C., J.S.K., W. Zhang; (LOLIPOP-OmniEE) J.C.C., J.S.K., Weihua Zhang; (MESA) J.I.R., X.G., J.Y.; (METSIM) X.S.; (Montreal Heart Institute Biobank (MHIBB)) J.C.T., G.L., K.S.L., V.T.; (OBB) A. Mahajan; (PCOS) A.P.M., T.K.; (PIVUS) N.R.R.; (PROMIS) A. Rasheed, W. Zhao; (quality control GoT2D/T2D-GENES (FUSION, METSIM and so on)) A.E.L.; (RISC) H.Y.; (Rotterdam Study I) C.M.G., F.R.; (SHIP/SHIP-Trend) A. Teumer; (SOLID TIMI-52) D.M.W., A.J.S.; (SORBS) A.P.M; (The Mount Sinai BioMe Biobank) Y.L., C.S.; (The NEO Study) R.L.G.; (The NHAPC study, The GBTDS study) X.L., H.L., Y.H., H.Z.; (The Western Australian Pregnancy Cohort (Raine) Study) C.A.W.; (UK Biobank) A.R.W.; (ULSAM) A.P.M., A. Mahajan; (WGHS) D.I.C., A.Y.C.; (Women’s Health Initiative) P.L.A., J.H.; (WTCCC-UKT2D) W.G.; (YFS) L.P.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interests: A.S.B. holds interest in AstraZeneca, Biogen, Bioverativ, Merck, Novartis and Pfizer. A.S.C. and C.S.F. are current employees of Merck. Authors affiliated with deCODE (V.St., G.T., U.T. and K.S.) are employed by deCODE Genetics/Amgen, Inc. H.D.W. has the following financial and non-financial competing interests to declare: research grants: Sanofi Aventis; Eli Lilly; NIH; Omthera Pharmaceuticals, Pfizer, Elsai Inc. AstraZeneca; DalCor and Services; lecture fees: Sanofi Aventis; advisory boards: Acetelion, Sirtex, CSL Boehring. J.D. has received grants from AstraZeneca, Biogen, Merck, Novartis and Pfizer. L.M.Y.A. and R.A.S. are employee stock holders of GlaxoSmithKline. M.P.D. received honoraria and holds minor equity in Dalcor. V.S. has participated in a conference trip sponsored by Novo Nordisk.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–24, Supplementary Tables 1–10 and Supplementary Note
Supplementary Data 1
Study design, number of individuals and sample quality control for ExomeChip study cohorts.
Supplementary Data 2
Information on genotyping methods, quality control of SNPs, imputation, and statistical analysis for ExomeChip study cohorts.
Supplementary Data 3
Study-specific descriptive statistics for anthropometric measurements in men and women of the ExomeChip cohorts.
Supplementary Data 4
Sex combined stage 1 and stage 2 results for all variants with initial stage 1 results that met P value
Supplementary Data 5
Male-Specific stage 1 and stage 2 results for all variants with initial stage 1 results that met P value
Supplementary Data 6
Female-Specific stage 1 and stage 2 results for all variants with initial stage 1 results that met P value
Supplementary Data 7
Conditional analysis and reciprocal
Supplementary Data 8
ExomeChip Data-driven ExpressionPrioritized Integration for Complex Traits (EC-DEPICT) results when including all variants associated WHRadjBMI with P value
Supplementary Data 9
ExomeChip Data-driven ExpressionPrioritized Integration for Complex Traits (EC-DEPICT) results when including all variants associated with WHRadjBMI with P value
Supplementary Data 10
Results from PASCAL (Pathway scoring algorithm) for Exomechip coding variants and genome-wide coding and regulatory variants associated with WHRadjBMI. Pascal (Pathway scoring algorithm) details and software are available at https://www2.unil.ch/cbg/index.php?title=Pascal
Supplementary Data 11
Associations between the WHRadjBMI that
Supplementary Data 12
Previously-reported associations with other traits in the GWAS Catalog for significant WHRadjBMI variants from the current study. This table lists all previously reported associations within 1 MB (+/- 500 kb) and in high LD (r2 >0.2) with our lead novel SNPs along with relevant annotation (for example miRNA target binding site, variant location relevant to nearest gene) reported in the GWAS Catalog.
Supplementary Data 13
Genes that have been linked to monogenic and syndromic obesity.
Supplementary Data 14
Exome variant - expression quantitative trait loci (eQTL) gene associations at false discovery rate
Supplementary Data 15
Detailed description for each of the variants that met array-wide significance in the stage 1 plus stage 2 sex-combined analyses, men only analyses, or women only analyses (see also Tables 1 and 2).
Supplementary Data 16
Exome variant - expression quantitative trait loci (eQTL) gene associations at false discovery rate <5% identified in the Genotype-Tissue Expression (GTEx) database. The results were extracted from the GTEx project (version 6):
Supplementary Data 17
Variant Effect Predictor (VEP) annotation
Supplementary Data 18
Detailed acknowledgements by study and/or contributing author(s).
Rights and permissions
About this article
Cite this article
Justice, A.E., Karaderi, T., Highland, H.M. et al. Protein-coding variants implicate novel genes related to lipid homeostasis contributing to body-fat distribution. Nat Genet 51, 452–469 (2019). https://doi.org/10.1038/s41588-018-0334-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0334-2
This article is cited by
-
SMYD3: a new regulator of adipocyte precursor proliferation at the early steps of differentiation
International Journal of Obesity (2024)
-
Genetics of sexually dimorphic adipose distribution in humans
Nature Genetics (2023)
-
Genetic regulation of body size and morphology in children: a twin study of 22 anthropometric traits
International Journal of Obesity (2023)
-
Genetics and epigenetics in the obesity phenotyping scenario
Reviews in Endocrine and Metabolic Disorders (2023)
-
Rare loss of function variants in the hepatokine gene INHBE protect from abdominal obesity
Nature Communications (2022)