-
 The Genome of the Yellow Mealworm, Tenebrio molitor: It’s Bigger Than You Think
The Genome of the Yellow Mealworm, Tenebrio molitor: It’s Bigger Than You Think -
 Microbiota-Induced Epigenetic Alterations in Depressive Disorders Are Targets for Nutritional and Probiotic Therapies
Microbiota-Induced Epigenetic Alterations in Depressive Disorders Are Targets for Nutritional and Probiotic Therapies -
 The Efficacy of a Human-Ready miniMECP2 Gene Therapy in a Pre-Clinical Model of Rett Syndrome
The Efficacy of a Human-Ready miniMECP2 Gene Therapy in a Pre-Clinical Model of Rett Syndrome -
 Pharmacogenomics of Dementia: Personalizing the Treatment of Cognitive and Neuropsychiatric Symptoms
Pharmacogenomics of Dementia: Personalizing the Treatment of Cognitive and Neuropsychiatric Symptoms
Journal Description
Genes
Genes
is a peer-reviewed, open access journal of genetics and genomics published monthly online by MDPI. The Spanish Society for Biochemistry and Molecular Biology (SEBBM) is affiliated with Genes and their members receive discounts on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, and other databases.
- Journal Rank: JCR - Q2 (Genetics & Heredity) / CiteScore - Q2 (Genetics)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.5 days after submission; acceptance to publication is undertaken in 2.3 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: Reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
3.5 (2022);
5-Year Impact Factor:
3.9 (2022)
Latest Articles
SNP and Structural Study of the Notch Superfamily Provides Insights and Novel Pharmacological Targets against the CADASIL Syndrome and Neurodegenerative Diseases
Genes 2024, 15(5), 529; https://doi.org/10.3390/genes15050529 - 23 Apr 2024
Abstract
The evolutionary conserved Notch signaling pathway functions as a mediator of direct cell–cell communication between neighboring cells during development. Notch plays a crucial role in various fundamental biological processes in a wide range of tissues. Accordingly, the aberrant signaling of this pathway underlies
[...] Read more.
The evolutionary conserved Notch signaling pathway functions as a mediator of direct cell–cell communication between neighboring cells during development. Notch plays a crucial role in various fundamental biological processes in a wide range of tissues. Accordingly, the aberrant signaling of this pathway underlies multiple genetic pathologies such as developmental syndromes, congenital disorders, neurodegenerative diseases, and cancer. Over the last two decades, significant data have shown that the Notch signaling pathway displays a significant function in the mature brains of vertebrates and invertebrates beyond neuronal development and specification during embryonic development. Neuronal connection, synaptic plasticity, learning, and memory appear to be regulated by this pathway. Specific mutations in human Notch family proteins have been linked to several neurodegenerative diseases including Alzheimer’s disease, CADASIL, and ischemic injury. Neurodegenerative diseases are incurable disorders of the central nervous system that cause the progressive degeneration and/or death of brain nerve cells, affecting both mental function and movement (ataxia). There is currently a lot of study being conducted to better understand the molecular mechanisms by which Notch plays an essential role in the mature brain. In this study, an in silico analysis of polymorphisms and mutations in human Notch family members that lead to neurodegenerative diseases was performed in order to investigate the correlations among Notch family proteins and neurodegenerative diseases. Particular emphasis was placed on the study of mutations in the Notch3 protein and the structure analysis of the mutant Notch3 protein that leads to the manifestation of the CADASIL syndrome in order to spot possible conserved mutations and interpret the effect of these mutations in the Notch3 protein structure. Conserved mutations of cysteine residues may be candidate pharmacological targets for the potential therapy of CADASIL syndrome.
Full article
(This article belongs to the Special Issue Head and Neck Genetics)
Open AccessArticle
Transcriptome Analysis on the Quality of Epimedium koreanum in Different Soil Moisture Conditions at Harvesting Stage
by
Yonggang Zhang, Dantong Wang, Feng Wu, Xiangdi Huang, Xiaowei Chai and Limin Yang
Genes 2024, 15(5), 528; https://doi.org/10.3390/genes15050528 - 23 Apr 2024
Abstract
Epimedium koreanum is a traditional Chinese tonic herb. Its main medicinal components are secondary metabolites such as flavonoids and flavonol glycosides, but the biosynthetic mechanism is still unclear. Moisture conditions are a key environmental factor affecting E. koreanum medicinal components during harvesting. Different
[...] Read more.
Epimedium koreanum is a traditional Chinese tonic herb. Its main medicinal components are secondary metabolites such as flavonoids and flavonol glycosides, but the biosynthetic mechanism is still unclear. Moisture conditions are a key environmental factor affecting E. koreanum medicinal components during harvesting. Different stages of E. koreanum under natural conditions after rainfall were selected to study changes in physiological properties, herb quality, and transcriptome. Malondialdehyde (MDA) content increased significantly in the D3 stage after rainfall, and protective enzyme levels also rose. Additionally, the flavonol glycoside content was relatively high. We sequenced the transcriptomes of D1, D3, and D9 (R) and identified differentially expressed genes (DEGs) related to flavonoid synthesis. This analysis allowed us to predict the roadmap and key genes involved in flavonoid biosynthesis for E. koreanum. These results suggest that the E. koreanum quality can be enhanced by natural drought conditions in the soil after precipitation during harvest. The harvesting period of E. koreanum is optimal when soil moisture naturally dries to a relative water content of 26% after precipitation. These conditions help E. koreanum tolerate a certain level of water scarcity, resulting in increased expression of flavonoid-related genes and ultimately enhancing the quality of the herb.
Full article
(This article belongs to the Section Plant Genetics and Genomics)
►▼
Show Figures
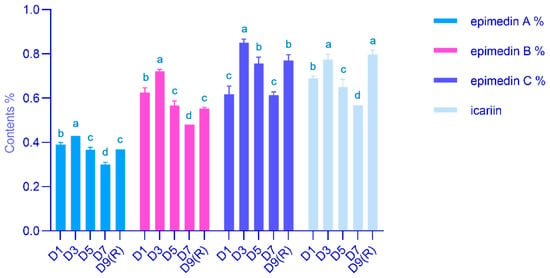
Figure 1
Open AccessArticle
Resolving Discrepancies in Idylla BRAF Mutational Assay Results Using Targeted Next-Generation Sequencing
by
Giby V. George, Huijie Liu, Audrey N. Jajosky and Zoltán N. Oltvai
Genes 2024, 15(5), 527; https://doi.org/10.3390/genes15050527 - 23 Apr 2024
Abstract
BRAF mutation identification is important for the diagnosis and treatment of several tumor types, both solid and hematologic. Rapid identification of BRAF mutations is required to determine eligibility for targeted BRAF inhibitor therapy. The Idylla BRAF mutation assay is a rapid, multiplex allele-specific
[...] Read more.
BRAF mutation identification is important for the diagnosis and treatment of several tumor types, both solid and hematologic. Rapid identification of BRAF mutations is required to determine eligibility for targeted BRAF inhibitor therapy. The Idylla BRAF mutation assay is a rapid, multiplex allele-specific PCR test designed to detect the most common oncogenic BRAF V600 mutations in formalin-fixed paraffin-embedded (FFPE) tissue samples. Here, we describe the validation of the Idylla BRAF mutation assay in our laboratory. During routine clinical practice, we noticed cases in which BRAF V600 mutations were identified with unusual amplification curves, with three cases displaying a delayed amplification within a double amplification pattern and two false-positive calls. We therefore initiated a quality improvement effort to systematically and retrospectively evaluate next-generation sequencing (NGS)-tested cases with BRAF mutations identified within five amino acids of BRAF codon V600 and did not identify additional false-positive cases. We hypothesize that late amplification in a double amplification pattern may represent non-specific amplification, whereas cases displaying single delayed amplification curves may stem from the presence of either non-V600 variants, very low-level V600 variants, cytosine deamination artifacts, and/or non-specific amplification by an allele-specific PCR primer. Regardless, we recommend that Idylla BRAF cases with non-classical amplification curves undergo reflex NGS testing. These findings are likely relevant for other Idylla assays interrogating hotspot mutations in genes such as EGFR, IDH1/2, KRAS, and NRAS.
Full article
(This article belongs to the Special Issue Precision Medicine and Genetics)
Open AccessArticle
Impact of Vanadium–Titanium–Magnetite Mining Activities on Endophytic Bacterial Communities and Functions in the Root Systems of Local Plants
by
Zhuang Xiong, Yunfeng Zhang, Xiaodie Chen, Ajia Sha, Wenqi Xiao, Yingyong Luo, Lianxin Peng, Liang Zou and Qiang Li
Genes 2024, 15(5), 526; https://doi.org/10.3390/genes15050526 - 23 Apr 2024
Abstract
This study utilized 16S rRNA high-throughput sequencing technology to analyze the community structure and function of endophytic bacteria within the roots of three plant species in the vanadium–titanium–magnetite (VTM) mining area. The findings indicated that mining activities of VTM led to a notable
[...] Read more.
This study utilized 16S rRNA high-throughput sequencing technology to analyze the community structure and function of endophytic bacteria within the roots of three plant species in the vanadium–titanium–magnetite (VTM) mining area. The findings indicated that mining activities of VTM led to a notable decrease in both the biodiversity and abundance of endophytic bacteria within the root systems of Eleusine indica and Carex (p < 0.05). Significant reductions were observed in the populations of Nocardioides, concurrently with substantial increments in the populations of Pseudomonas (p < 0.05), indicating that Pseudomonas has a strong adaptability to this environmental stress. In addition, β diversity analysis revealed divergence in the endophytic bacterial communities within the roots of E. indica and Carex from the VTM mining area, which had diverged to adapt to the environmental stress caused by mining activity. Functional enrichment analysis revealed that VTM mining led to an increase in polymyxin resistance, nicotinate degradation I, and glucose degradation (oxidative) (p < 0.05). Interestingly, we found that VTM mining did not notably alter the endophytic bacterial communities or functions in the root systems of Dodonaea viscosa, indicating that this plant can adapt well to environmental stress. This study represents the primary investigation into the influence of VTM mining activities on endophytic bacterial communities and the functions of nearby plant roots, providing further insight into the impact of VTM mining activities on the ecological environment.
Full article
(This article belongs to the Special Issue Genomics of Microbial Diversity, Evolution and Function)
Open AccessCase Report
A TMEM63A Nonsense Heterozygous Variant Linked to Infantile Transient Hypomyelinating Leukodystrophy Type 19?
by
Dimitra Siori, Dimitrios Vlachakis, Periklis Makrythanasis, Joanne Traeger-Synodinos, Danai Veltra, Afrodite Kampouraki and George P. Chrousos
Genes 2024, 15(5), 525; https://doi.org/10.3390/genes15050525 - 23 Apr 2024
Abstract
Infantile onset transient hypomyelination (IOTH) is a rare form of leukodystrophy that is associated with transient motor impairment and delayed central nervous system myelination. Here, we report a case of a new mutation in the transmembrane protein 63A (TMEM63A) gene identified
[...] Read more.
Infantile onset transient hypomyelination (IOTH) is a rare form of leukodystrophy that is associated with transient motor impairment and delayed central nervous system myelination. Here, we report a case of a new mutation in the transmembrane protein 63A (TMEM63A) gene identified using Whole-Exome Sequencing (WES) in an 8.5-year-old boy with clinical symptoms similar to IOTH. The patient exhibited a mild developmental delay, including hypotonia and delayed motor milestones, as well as some notable phenotypic characteristics, such as macrocephaly and macrosomia. Despite the absence of early neuroimaging, genetic testing revealed a paternally inherited variant in TMEM63A (NM_14698.3:c.220A>T;p:(Arg74*)), potentially linked to infantile transient hypomyelinating leukodystrophy type 19. Our findings in this study and the patient’s favorable clinical course underscore the potential for successful myelination even with delayed initiation and may contribute to a better understanding of the genotype–phenotype correlation in IOTH, emphasizing the importance of genetic analysis in unresolved developmental delay cases and providing critical insights for accurate diagnosis, prognosis and potential therapeutic strategies in rare leukodystrophies.
Full article
(This article belongs to the Special Issue Head and Neck Genetics)
►▼
Show Figures
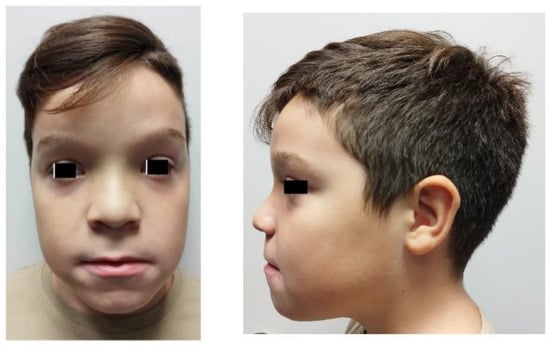
Figure 1
Open AccessArticle
Dissecting Selective Signatures and Candidate Genes in Grandparent Lines Subject to High Selection Pressure for Broiler Production and in a Local Russian Chicken Breed of Ushanka
by
Michael N. Romanov, Alexey V. Shakhin, Alexandra S. Abdelmanova, Natalia A. Volkova, Dmitry N. Efimov, Vladimir I. Fisinin, Liudmila G. Korshunova, Dmitry V. Anshakov, Arsen V. Dotsev, Darren K. Griffin and Natalia A. Zinovieva
Genes 2024, 15(4), 524; https://doi.org/10.3390/genes15040524 - 22 Apr 2024
Abstract
Breeding improvements and quantitative trait genetics are essential to the advancement of broiler production. The impact of artificial selection on genomic architecture and the genetic markers sought remains a key area of research. Here, we used whole-genome resequencing data to analyze the genomic
[...] Read more.
Breeding improvements and quantitative trait genetics are essential to the advancement of broiler production. The impact of artificial selection on genomic architecture and the genetic markers sought remains a key area of research. Here, we used whole-genome resequencing data to analyze the genomic architecture, diversity, and selective sweeps in Cornish White (CRW) and Plymouth Rock White (PRW) transboundary breeds selected for meat production and, comparatively, in an aboriginal Russian breed of Ushanka (USH). Reads were aligned to the reference genome bGalGal1.mat.broiler.GRCg7b and filtered to remove PCR duplicates and low-quality reads using BWA-MEM2 and bcftools software; 12,563,892 SNPs were produced for subsequent analyses. Compared to CRW and PRW, USH had a lower diversity and a higher genetic distinctiveness. Selective sweep regions and corresponding candidate genes were examined based on ZFST, hapFLK, and ROH assessment procedures. Twenty-seven prioritized chicken genes and the functional projection from human homologs suggest their importance for selection signals in the studied breeds. These genes have a functional relationship with such trait categories as body weight, muscles, fat metabolism and deposition, reproduction, etc., mainly aligned with the QTLs in the sweep regions. This information is pivotal for further executing genomic selection to enhance phenotypic traits.
Full article
(This article belongs to the Special Issue Poultry Genetics and Genomics (Volume II))
Open AccessArticle
Effects of Long-Term Cryopreservation on the Transcriptomes of Giant Grouper Sperm
by
Xiaoyu Ding, Yongsheng Tian, Yishu Qiu, Pengfei Duan, Xinyi Wang, Zhentong Li, Linlin Li, Yang Liu and Linna Wang
Genes 2024, 15(4), 523; https://doi.org/10.3390/genes15040523 - 22 Apr 2024
Abstract
The giant grouper fish (Epinephelus lanceolatus), one of the largest and rarest groupers, is a fast-growing economic fish. Grouper sperm is often used for cross-breeding with other fish and therefore sperm cryopreservation is important. However, freezing damage cannot be avoided. Herein,
[...] Read more.
The giant grouper fish (Epinephelus lanceolatus), one of the largest and rarest groupers, is a fast-growing economic fish. Grouper sperm is often used for cross-breeding with other fish and therefore sperm cryopreservation is important. However, freezing damage cannot be avoided. Herein, we performed a transcriptome analysis to compare fresh and frozen sperm of the giant grouper with frozen storage times of 0, 23, 49, and 61 months. In total, 1911 differentially expressed genes (DEGs), including 91 in El-0-vs-El-23 (40 upregulated and 51 downregulated), 251 in El-0-vs-El-49 (152 upregulated and 69 downregulated), and 1569 in El-0-vs-El-61 (984 upregulated and 585 downregulated), were obtained in the giant grouper sperm. DEGs were significantly increased at 61 months of cryopreservation (p < 0.05). GO and KEGG enrichment analyses of the DEGs revealed significant enrichment in the pilus assembly, metabolic process, MAPK signaling pathway, apoptosis, and P53 signaling pathway. Time-series expression profiling of the DEGs showed that consistently upregulated modules were also significantly enriched in signaling pathways associated with apoptosis. Four genes, scarb1, odf3, exoc8, and atp5f1d, were associated with mitochondria and flagella in a weighted correlation network analysis. These genes may play an important role in the response to sperm freezing. The experimental results show that long-term cryopreservation results in freezing damage to the giant grouper sperm. This study provides rich data for studies of the mechanism underlying frozen fish sperm damage as well as a technical reference and evaluation index for the long-term cryopreservation of fish sperm.
Full article
(This article belongs to the Special Issue Genetic Studies of Fish)
►▼
Show Figures
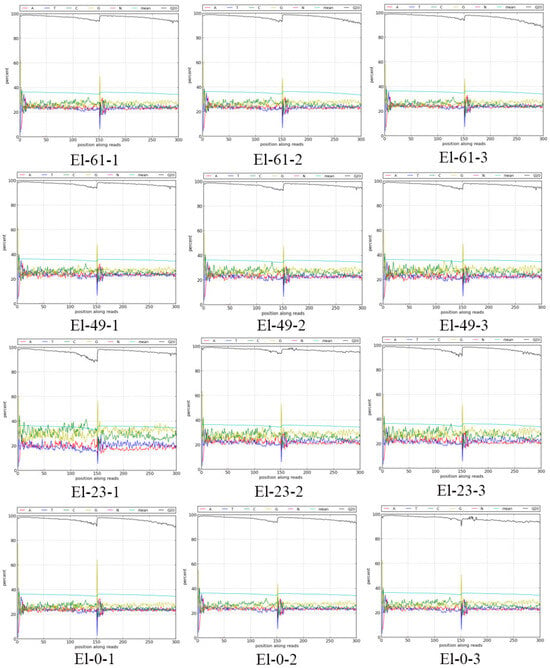
Figure 1
Open AccessArticle
HLA-B and C Expression Contributes to COVID-19 Disease Severity within a South African Cohort
by
Lisa Naidoo, Thilona Arumugam and Veron Ramsuran
Genes 2024, 15(4), 522; https://doi.org/10.3390/genes15040522 - 22 Apr 2024
Abstract
Globally, SARS-CoV-2 has negatively impacted many lives and industries due to its rapid spread, severe outcomes, and the need for the implementation of lockdown strategies across the world. SARS-CoV-2 disease severity varies among different populations. Host genetics have been associated with various diseases,
[...] Read more.
Globally, SARS-CoV-2 has negatively impacted many lives and industries due to its rapid spread, severe outcomes, and the need for the implementation of lockdown strategies across the world. SARS-CoV-2 disease severity varies among different populations. Host genetics have been associated with various diseases, and their ability to alter disease susceptibility and severity. In addition, Human Leukocyte Antigen (HLA) expression levels and alleles vary significantly among ethnic groups, which might impact the host’s response to SARS-CoV-2. Our previous study highlighted that HLA-A might have an effect on COVID-19 disease severity across ethnicities. Therefore, in this study, we aim to examine the effect of HLA-B and C expression levels on COVID-19 disease severity. To achieve this, we used real-time PCR to measure the HLA mRNA expression levels of SARS-CoV-2-infected individuals from a South African cohort and compared them across ethnic groups, disease outcomes, gender, comorbidities, and age. Our results show (1) that the effect of HLA-B mRNA expression levels was associated with differences in disease severity when we compare symptomatic vs. asymptomatic (p < 0.0001). While HLA-C mRNA expression levels were not associated with COVID-19 disease severity. (2) In addition, we observed that HLA-B and HLA-C mRNA expression levels were significantly different between South African Black individuals and South African Indian individuals (p < 0.0001, p < 0.0001). HLA-B mRNA expression levels among symptomatic South African Black individuals were significantly higher than symptomatic South African Indian individuals (p < 0.0001). In addition, the HLA-B mRNA expression levels of symptomatic South African Black individuals were significantly higher than asymptomatic South African Black individuals (p > 0.0001). HLA-C mRNA expression levels among symptomatic South African Black individuals were significantly higher than among symptomatic South African Indian individuals (p = 0.0217). (3) HLA-C expression levels were significantly different between males and females (p = 0.0052). In addition, asymptomatic males are higher than asymptomatic females (p = 0.0375). (4) HLA-B expression levels were significantly different between individuals with and without comorbidities (p = 0.0009). In addition, we observed a significant difference between individuals with no comorbidities and non-communicable diseases (p = 0.0034), in particular, hypertension (p = 0.0487). (5) HLA-B expression levels were significantly different between individuals between 26–35 and 56–65 years (p = 0.0380). Our work is expected to strengthen the understanding of the relationship between HLA and COVID-19 by providing insights into HLA-B and C expression levels across ethnic populations in South Africa among COVID-19-symptomatic and asymptomatic individuals. Our results highlight that HLA-B mRNA expression levels contribute to COVID-19 severity as well as variation in ethnicities associated with COVID-19. Further studies are needed to examine the effect of HLA expression levels across various ethnic groups with contributing factors.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
►▼
Show Figures
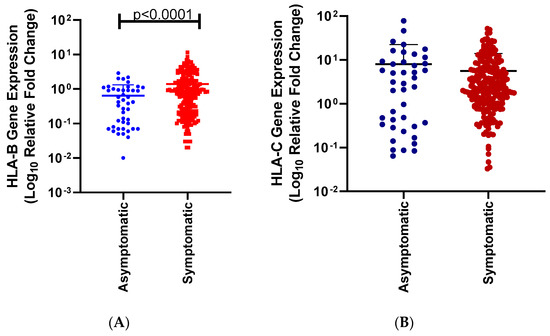
Figure 1
Open AccessArticle
Contradiction in Star-Allele Nomenclature of Pharmacogenes between Common Haplotypes and Rare Variants
by
Se Hwan Ahn, Yoomi Park and Ju Han Kim
Genes 2024, 15(4), 521; https://doi.org/10.3390/genes15040521 - 22 Apr 2024
Abstract
The nomenclature of star alleles has been widely used in pharmacogenomics to enhance treatment outcomes, predict drug response variability, and reduce adverse reactions. However, the discovery of numerous rare functional variants through genome sequencing introduces complexities into the star-allele system. This study aimed
[...] Read more.
The nomenclature of star alleles has been widely used in pharmacogenomics to enhance treatment outcomes, predict drug response variability, and reduce adverse reactions. However, the discovery of numerous rare functional variants through genome sequencing introduces complexities into the star-allele system. This study aimed to assess the nature and impact of the rapid discovery of numerous rare functional variants in the traditional haplotype-based star-allele system. We developed a new method to construct haplogroups, representing a common ancestry structure, by iteratively excluding rare and functional variants of the 25 representative pharmacogenes using the 2504 genomes from the 1000 Genomes Project. In total, 192 haplogroups and 288 star alleles were identified, with an average of 7.68 ± 4.2 cross-ethnic haplogroups per gene. Most of the haplogroups (70.8%, 136/192) were highly aligned with their corresponding classical star alleles (VI = 1.86 ± 0.78), exhibiting higher genetic diversity than the star alleles. Approximately 41.3% (N = 119) of the star alleles in the 2504 genomes did not belong to any of the haplogroups, and most of them (91.3%, 105/116) were determined by a single variant according to the allele-definition table provided by CPIC. These functional single variants had low allele frequency (MAF < 1%), high evolutionary conservation, and variant deleteriousness, which suggests significant negative selection. It is suggested that the traditional haplotype-based naming system for pharmacogenetic star alleles now needs to be adjusted by balancing both traditional haplotyping and newly emerging variant-sequencing approaches to reduce naming complexity.
Full article
(This article belongs to the Section Pharmacogenetics)
►▼
Show Figures

Figure 1
Open AccessCommentary
DNA Damage, Genome Stability, and Adaptation: A Question of Chance or Necessity?
by
John Herrick
Genes 2024, 15(4), 520; https://doi.org/10.3390/genes15040520 - 21 Apr 2024
Abstract
DNA damage causes the mutations that are the principal source of genetic variation. DNA damage detection and repair mechanisms therefore play a determining role in generating the genetic diversity on which natural selection acts. Speciation, it is commonly assumed, occurs at a rate
[...] Read more.
DNA damage causes the mutations that are the principal source of genetic variation. DNA damage detection and repair mechanisms therefore play a determining role in generating the genetic diversity on which natural selection acts. Speciation, it is commonly assumed, occurs at a rate set by the level of standing allelic diversity in a population. The process of speciation is driven by a combination of two evolutionary forces: genetic drift and ecological selection. Genetic drift takes place under the conditions of relaxed selection, and results in a balance between the rates of mutation and the rates of genetic substitution. These two processes, drift and selection, are necessarily mediated by a variety of mechanisms guaranteeing genome stability in any given species. One of the outstanding questions in evolutionary biology concerns the origin of the widely varying phylogenetic distribution of biodiversity across the Tree of Life and how the forces of drift and selection contribute to shaping that distribution. The following examines some of the molecular mechanisms underlying genome stability and the adaptive radiations that are associated with biodiversity and the widely varying species richness and evenness in the different eukaryotic lineages.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
►▼
Show Figures
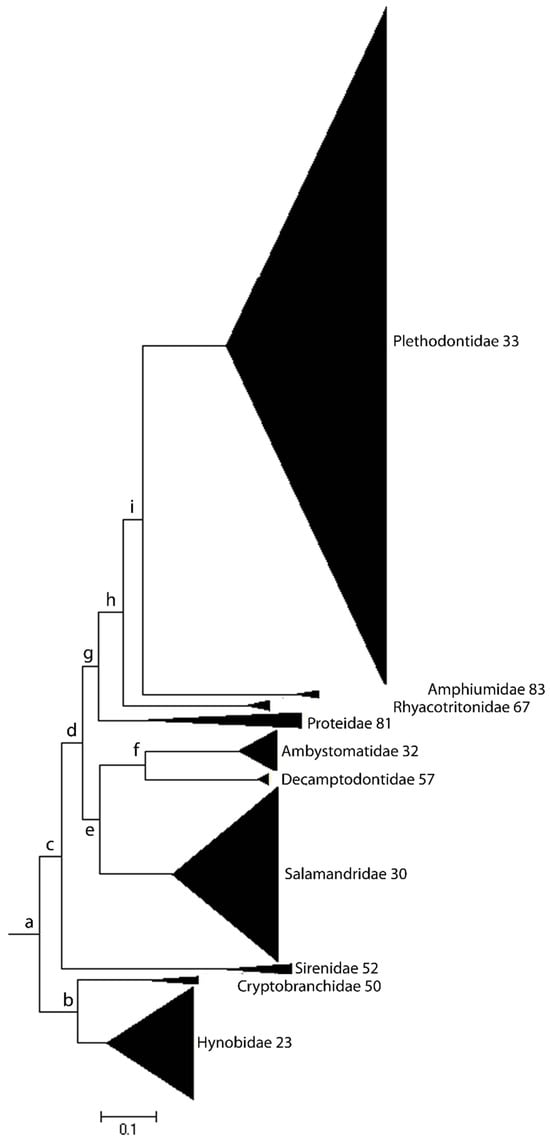
Figure 1
Open AccessArticle
Single-Nucleus Transcriptome Profiling from the Hippocampus of a PTSD Mouse Model and CBD-Treated Cohorts
by
Guanbo Xie, Yihan Qin, Ning Wu, Xiao Han and Jin Li
Genes 2024, 15(4), 519; https://doi.org/10.3390/genes15040519 - 21 Apr 2024
Abstract
Post-traumatic stress disorder (PTSD) is the most common psychiatric disorder after a catastrophic event; however, the efficacious treatment options remain insufficient. Increasing evidence suggests that cannabidiol (CBD) exhibits optimal therapeutic effects for treating PTSD. To elucidate the cell-type-specific transcriptomic pathology of PTSD and
[...] Read more.
Post-traumatic stress disorder (PTSD) is the most common psychiatric disorder after a catastrophic event; however, the efficacious treatment options remain insufficient. Increasing evidence suggests that cannabidiol (CBD) exhibits optimal therapeutic effects for treating PTSD. To elucidate the cell-type-specific transcriptomic pathology of PTSD and the mechanisms of CBD against this disease, we conducted single-nucleus RNA sequencing (snRNA-seq) in the hippocampus of PTSD-modeled mice and CBD-treated cohorts. We constructed a mouse model by adding electric foot shocks following exposure to single prolonged stress (SPS+S) and tested the freezing time, anxiety-like behavior, and cognitive behavior. CBD was administrated before every behavioral test. The PTSD-modeled mice displayed behaviors resembling those of PTSD in all behavioral tests, and CBD treatment alleviated all of these PTSD-like behaviors (n = 8/group). Three mice with representative behavioral phenotypes were selected from each group for snRNA-seq 15 days after the SPS+S. We primarily focused on the excitatory neurons (ExNs) and inhibitory neurons (InNs), which accounted for 68.4% of the total cell annotations. A total of 88 differentially upregulated genes and 305 differentially downregulated genes were found in the PTSD mice, which were found to exhibit significant alterations in pathways and biological processes associated with fear response, synaptic communication, protein synthesis, oxidative phosphorylation, and oxidative stress response. A total of 63 overlapping genes in InNs were identified as key genes for CBD in the treatment of PTSD. Subsequent Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses revealed that the anti-PTSD effect of CBD was related to the regulation of protein synthesis, oxidative phosphorylation, oxidative stress response, and fear response. Furthermore, gene set enrichment analysis (GSEA) revealed that CBD also enhanced retrograde endocannabinoid signaling in ExNs, which was found to be suppressed in the PTSD group. Our research may provide a potential explanation for the pathogenesis of PTSD and facilitate the discovery of novel therapeutic targets for drug development. Moreover, it may shed light on the therapeutic mechanisms of CBD.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
►▼
Show Figures
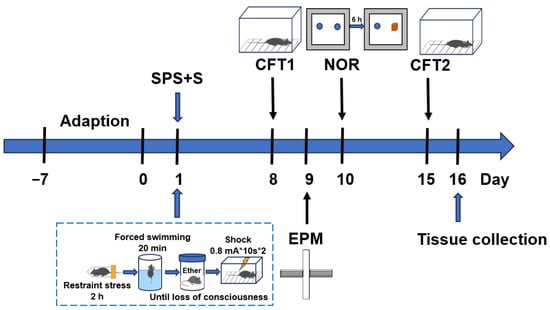
Figure 1
Open AccessArticle
22q11.2 Deletion Syndrome: Influence of Parental Origin on Clinical Heterogeneity
by
Melissa Bittencourt de Wallau, Ana Carolina Xavier, Carolina Araújo Moreno, Chong Ae Kim, Elaine Lustosa Mendes, Erlane Marques Ribeiro, Amanda Oliveira, Têmis Maria Félix, Agnes Cristina Fett-Conte, Luciana Cardoso Bonadia, Gabriela Roldão Correia-Costa, Isabella Lopes Monlleó, Vera Lúcia Gil-da-Silva-Lopes and Társis Paiva Vieira
Genes 2024, 15(4), 518; https://doi.org/10.3390/genes15040518 - 21 Apr 2024
Abstract
22q11.2 deletion syndrome (22q11.2DS) shows significant clinical heterogeneity. This study aimed to explore the association between clinical heterogeneity in 22q11.2DS and the parental origin of the deletion. The parental origin of the deletion was determined for 61 individuals with 22q11.2DS by genotyping DNA
[...] Read more.
22q11.2 deletion syndrome (22q11.2DS) shows significant clinical heterogeneity. This study aimed to explore the association between clinical heterogeneity in 22q11.2DS and the parental origin of the deletion. The parental origin of the deletion was determined for 61 individuals with 22q11.2DS by genotyping DNA microsatellite markers and single-nucleotide polymorphisms (SNPs). Among the 61 individuals, 29 (47.5%) had a maternal origin of the deletion, and 32 (52.5%) a paternal origin. Comparison of the frequency of the main clinical features between individuals with deletions of maternal or paternal origin showed no statistically significant difference. However, Truncus arteriosus, pulmonary atresia, seizures, and scoliosis were only found in patients with deletions of maternal origin. Also, a slight difference in the frequency of other clinical features between groups of maternal or paternal origin was noted, including congenital heart disease, endocrinological alterations, and genitourinary abnormalities, all of them more common in patients with deletions of maternal origin. Although parental origin of the deletion does not seem to contribute to the phenotypic variability of most clinical signs observed in 22q11.2DS, these findings suggest that patients with deletions of maternal origin could have a more severe phenotype. Further studies with larger samples focusing on these specific features could corroborate these findings.
Full article
(This article belongs to the Special Issue Phenotype and Pathogenetic Mechanisms in 22q11.2 Deletion/DiGeorge Syndrome)
►▼
Show Figures
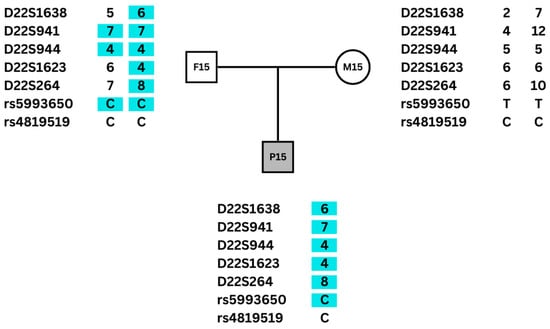
Figure 1
Open AccessArticle
Information Provision Regarding Health-Related Direct-to-Consumer Genetic Testing for Dutch Consumers: An in-Depth Content Analysis of Sellers’ Websites
by
Danny Bruins, Suzanne M. Onstwedder, Martina C. Cornel, Margreet G. E. M. Ausems, Marc H. W. van Mil and Tessel Rigter
Genes 2024, 15(4), 517; https://doi.org/10.3390/genes15040517 - 20 Apr 2024
Abstract
Background: Previous studies have suggested that information offered by sellers of health-related direct-to-consumer genetic tests (DTC-GTs) is often incomplete, unbalanced, or too difficult to understand. The extent to which this is the case for sellers accessible to Dutch consumers has not previously
[...] Read more.
Background: Previous studies have suggested that information offered by sellers of health-related direct-to-consumer genetic tests (DTC-GTs) is often incomplete, unbalanced, or too difficult to understand. The extent to which this is the case for sellers accessible to Dutch consumers has not previously been studied. Methods and Goals: The present study aimed to assess the completeness, balance, readability, and findability of informational content on a selection of websites from several health-related DTC-GT sellers accessible to Dutch consumers. An in-depth content analysis was performed based on a recently published checklist outlining key items for policy guidance regarding DTC-GT services. Results: The information provided by sellers did not equally cover all aspects relevant to health-related DTC-GT service provision. The provided information was slightly unbalanced, with benefits of health-related DTC-GT usage being overemphasized compared to its risks and limitations. The readability of the provided information was low, on average requiring college education for proper understanding. A findability analysis showed that information concerning all themes is overall relatively evenly distributed across analyzed sellers’ websites. Conclusions: Information provision by assessed health-related DTC-GT sellers is suboptimal regarding completeness, balance, and readability. To better empower potential consumers to make an informed decision regarding health-related DTC-GT usage, we advocate industry-wide enhancement of information provision.
Full article
(This article belongs to the Special Issue Human Genetics: Diseases, Community, and Counseling)
►▼
Show Figures
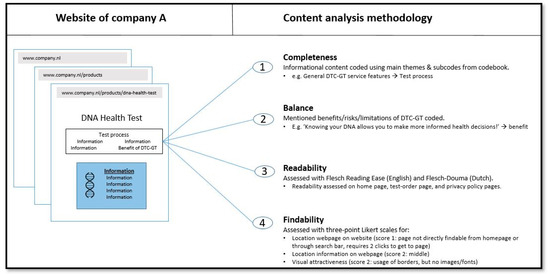
Figure 1
Open AccessCase Report
Syndromic Retinitis Pigmentosa: A 15-Patient Study
by
Ianne Pessoa Holanda, Priscila Hae Hyun Rim, Rare Genomes Project Consortium, Mara Sanches Guaragna, Vera Lúcia Gil-da-Silva-Lopes and Carlos Eduardo Steiner
Genes 2024, 15(4), 516; https://doi.org/10.3390/genes15040516 - 20 Apr 2024
Abstract
Retinitis pigmentosa is a group of genetically determined retinal dystrophies characterized by primary photoreceptor apoptosis and can occur in isolated or syndromic conditions. This study reviewed the clinical data of 15 patients with syndromic retinitis pigmentosa from a Rare Disease Reference Center in
[...] Read more.
Retinitis pigmentosa is a group of genetically determined retinal dystrophies characterized by primary photoreceptor apoptosis and can occur in isolated or syndromic conditions. This study reviewed the clinical data of 15 patients with syndromic retinitis pigmentosa from a Rare Disease Reference Center in Brazil and the results of their next-generation sequencing tests. Five males and ten females participated, with the mean ages for ocular disease onset, fundoscopic diagnosis, and molecular evaluation being 9, 19, and 29 years, respectively. Bardet–Biedl syndrome (n = 5) and Usher syndrome (n = 3) were the most frequent diagnoses, followed by other rare conditions. Among the patients, fourteen completed molecular studies, with three negative results and eleven revealing findings in known genes, including novel variants in MKKS (c.432_435del, p.Phe144Leufs*14), USH2A (c.(7301+1_7302-1)_(9369+1_9370-1)del), and CEP250 (c.5383dup, p.Glu1795Glyfs*13, and c.5050del, p.Asp1684Thrfs*9). Except for Kearn-Sayre, all presented an autosomal recessive inheritance pattern with 64% homozygosity results. The long gap between symptom onset and diagnosis highlights the diagnostic challenges faced by the patients. This study reaffirms the clinical heterogeneity of syndromic retinitis pigmentosa and underscores the pivotal role of molecular analysis in advancing our understanding of these diseases.
Full article
(This article belongs to the Special Issue Molecular Diagnosis and Disease Mechanisms in Eye Disorders)
►▼
Show Figures

Figure 1
Open AccessArticle
Identification of Crucial Modules and Genes Associated with Bt Gene Expression in Cotton
by
Guiyuan Zhao, Zhao Geng, Jianguang Liu, Haiyan Tian, Xu Liu, Zetong An, Ning Zhao, Hanshuang Zhang, Liqiang Wu, Xingfen Wang, Yongqiang Wang and Guiyin Zhang
Genes 2024, 15(4), 515; https://doi.org/10.3390/genes15040515 - 19 Apr 2024
Abstract
The expression of Bacillus thuringiensis (Bt) toxins in transgenic cotton confers resistance to insect pests. However, it has been demonstrated that its effectiveness varies among cotton cultivars and different tissues. In this study, we evaluated the expression of Bt protein in 28 cotton
[...] Read more.
The expression of Bacillus thuringiensis (Bt) toxins in transgenic cotton confers resistance to insect pests. However, it has been demonstrated that its effectiveness varies among cotton cultivars and different tissues. In this study, we evaluated the expression of Bt protein in 28 cotton cultivars and selected 7 cultivars that differed in Bt protein expression for transcriptome analysis. Based on their Bt protein expression levels, the selected cultivars were categorized into three groups: H (high Bt protein expression), M (moderate expression), and L (low expression). In total, 342, 318, and 965 differentially expressed genes were detected in the H vs. L, M vs. L, and H vs. M comparison groups, respectively. And three modules significantly associated with Bt protein expression were identified by weighted gene co-expression network analysis. Three hub genes were selected to verify their relationships with Bt protein expression using virus-induced gene silencing (VIGS). Silencing GhM_D11G1176, encoding an MYC transcription factor, was confirmed to significantly decrease the expression of Bt protein. The present findings contribute to an improved understanding of the mechanisms that influence Bt protein expression in transgenic cotton.
Full article
(This article belongs to the Special Issue Cotton Genes, Genetics, and Genomics)
►▼
Show Figures
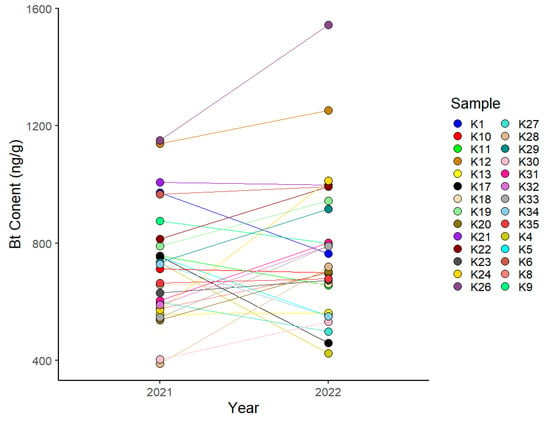
Figure 1
Open AccessArticle
A High-Quality Assembly and Comparative Analysis of the Mitogenome of Actinidia macrosperma
by
Jiangmei Gong, Jun Yang, Yan Lai, Tengfei Pan and Wenqin She
Genes 2024, 15(4), 514; https://doi.org/10.3390/genes15040514 - 19 Apr 2024
Abstract
The mitochondrial genome (mitogenome) of Actinidia macrosperma, a traditional medicinal plant within the Actinidia genus, remains relatively understudied. This study aimed to sequence the mitogenome of A. macrosperma, determining its assembly, informational content, and developmental expression. The results revealed that the
[...] Read more.
The mitochondrial genome (mitogenome) of Actinidia macrosperma, a traditional medicinal plant within the Actinidia genus, remains relatively understudied. This study aimed to sequence the mitogenome of A. macrosperma, determining its assembly, informational content, and developmental expression. The results revealed that the mitogenome of A. macrosperma is circular, spanning 752,501 bp with a GC content of 46.16%. It comprises 63 unique genes, including 39 protein-coding genes (PCGs), 23 tRNA genes, and three rRNA genes. Moreover, the mitogenome was found to contain 63 SSRs, predominantly mono-nucleotides, as well as 25 tandem repeats and 650 pairs of dispersed repeats, each with lengths equal to or greater than 60, mainly comprising forward repeats and palindromic repeats. Moreover, 53 homologous fragments were identified between the mitogenome and chloroplast genome (cp-genome), with the longest segment measuring 4296 bp. This study represents the initial report on the mitogenome of the A. macrosperma, providing crucial genetic materials for phylogenetic research within the Actinidia genus and promoting the exploitation of species genetic resources.
Full article
(This article belongs to the Section Plant Genetics and Genomics)
►▼
Show Figures

Figure 1
Open AccessCase Report
Genome Sequencing in an Individual Presenting with 22q11.2 Deletion Syndrome and Juvenile Idiopathic Arthritis
by
Ruy Pires de Oliveira-Sobrinho, Simone Appenzeller, Ianne Pessoa Holanda, Júlia Lôndero Heleno, Josep Jorente, on behalf of the Rare Genomes Project Consortium, Társis Paiva Vieira and Carlos Eduardo Steiner
Genes 2024, 15(4), 513; https://doi.org/10.3390/genes15040513 - 19 Apr 2024
Abstract
Juvenile idiopathic arthritis is a heterogeneous group of diseases characterized by arthritis with poorly known causes, including monogenic disorders and multifactorial etiology. 22q11.2 proximal deletion syndrome is a multisystemic disease with over 180 manifestations already described. In this report, the authors describe a
[...] Read more.
Juvenile idiopathic arthritis is a heterogeneous group of diseases characterized by arthritis with poorly known causes, including monogenic disorders and multifactorial etiology. 22q11.2 proximal deletion syndrome is a multisystemic disease with over 180 manifestations already described. In this report, the authors describe a patient presenting with a short stature, neurodevelopmental delay, and dysmorphisms, who had an episode of polyarticular arthritis at the age of three years and eight months, resulting in severe joint limitations, and was later diagnosed with 22q11.2 deletion syndrome. Investigation through Whole Genome Sequencing revealed that he had no pathogenic or likely-pathogenic variants in both alleles of the MIF gene or in genes associated with monogenic arthritis (LACC1, LPIN2, MAFB, NFIL3, NOD2, PRG4, PRF1, STX11, TNFAIP3, TRHR, UNC13DI). However, the patient presented 41 risk polymorphisms for juvenile idiopathic arthritis. Thus, in the present case, arthritis seems coincidental to 22q11.2 deletion syndrome, probably caused by a multifactorial etiology. The association of the MIF gene in individuals previously described with juvenile idiopathic arthritis and 22q11.2 deletion seems unlikely since it is located in the distal and less-frequently deleted region of 22q11.2 deletion syndrome.
Full article
(This article belongs to the Special Issue Phenotype and Pathogenetic Mechanisms in 22q11.2 Deletion/DiGeorge Syndrome)
►▼
Show Figures
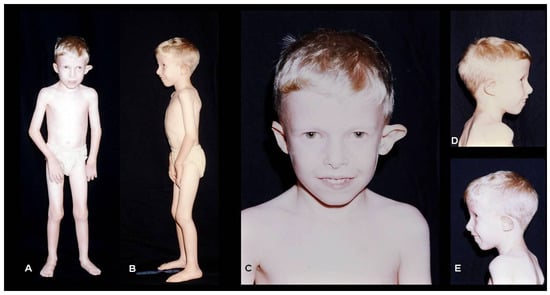
Figure 1
Open AccessArticle
Association between MCU Gene Polymorphisms with Obesity: Findings from the All of Us Research Program
by
Jade Avery, Tennille Leak-Johnson and Sharon C. Francis
Genes 2024, 15(4), 512; https://doi.org/10.3390/genes15040512 - 19 Apr 2024
Abstract
Obesity is a public health crisis, and its prevalence disproportionately affects African Americans in the United States. Dysregulation of organelle calcium homeostasis is associated with obesity. The mitochondrial calcium uniporter (MCU) complex is primarily responsible for mitochondrial calcium homeostasis. Obesity is
[...] Read more.
Obesity is a public health crisis, and its prevalence disproportionately affects African Americans in the United States. Dysregulation of organelle calcium homeostasis is associated with obesity. The mitochondrial calcium uniporter (MCU) complex is primarily responsible for mitochondrial calcium homeostasis. Obesity is a multifactorial disease in which genetic underpinnings such as single-nucleotide polymorphisms (SNPs) may contribute to disease progression. The objective of this study was to identify genetic variations of MCU with anthropometric measurements and obesity in the All of Us Research Program. Methods: We used an additive genetic model to assess the association between obesity traits (body mass index (BMI), waist and hip circumference) and selected MCU SNPs in 19,325 participants (3221 normal weight and 16,104 obese). Eleven common MCU SNPs with a minor allele frequency ≥ 5% were used for analysis. Results: We observed three MCU SNPs in self-reported Black/African American (B/AA) men, and six MCU SNPs in B/AA women associated with increased risk of obesity, whereas six MCU SNPs in White men, and nine MCU SNPs in White women were protective against obesity development. Conclusions: This study found associations of MCU SNPs with obesity, providing evidence of a potential predictor of obesity susceptibility in B/AA adults.
Full article
(This article belongs to the Special Issue Genetics of Obesity)
Open AccessArticle
An R2R3-MYB Transcriptional Factor LuMYB314 Associated with the Loss of Petal Pigmentation in Flax (Linum usitatissimum L.)
by
Dongliang Guo, Haixia Jiang and Liqiong Xie
Genes 2024, 15(4), 511; https://doi.org/10.3390/genes15040511 - 18 Apr 2024
Abstract
The loss of anthocyanin pigments is one of the most common evolutionary transitions in petal color, yet the genetic basis for these changes in flax remains largely unknown. In this study, we used crossing studies, a bulk segregant analysis, genome-wide association studies, a
[...] Read more.
The loss of anthocyanin pigments is one of the most common evolutionary transitions in petal color, yet the genetic basis for these changes in flax remains largely unknown. In this study, we used crossing studies, a bulk segregant analysis, genome-wide association studies, a phylogenetic analysis, and transgenic testing to identify genes responsible for the transition from blue to white petals in flax. This study found no correspondence between the petal color and seed color, refuting the conclusion that a locus controlling the seed coat color is associated with the petal color, as reported in previous studies. The locus controlling the petal color was mapped using a BSA-seq analysis based on the F2 population. However, no significantly associated genomic regions were detected. Our genome-wide association study identified a highly significant QTL (BP4.1) on chromosome 4 associated with flax petal color in the natural population. The combination of a local Manhattan plot and an LD heat map identified LuMYB314, an R2R3-MYB transcription factor, as a potential gene responsible for the natural variations in petal color in flax. The overexpression of LuMYB314 in both Arabidopsis thaliana and Nicotiana tabacum resulted in anthocyanin deposition, indicating that LuMYB314 is a credible candidate gene for controlling the petal color in flax. Additionally, our study highlights the limitations of the BSA-seq method in low-linkage genomic regions, while also demonstrating the powerful detection capabilities of GWAS based on high-density genomic variation mapping. This study enhances our genetic insight into petal color variations and has potential breeding value for engineering LuMYB314 to develop colored petals, bast fibers, and seeds for multifunctional use in flax.
Full article
(This article belongs to the Special Issue Advances in Genetics and Genomics of Plants)
►▼
Show Figures
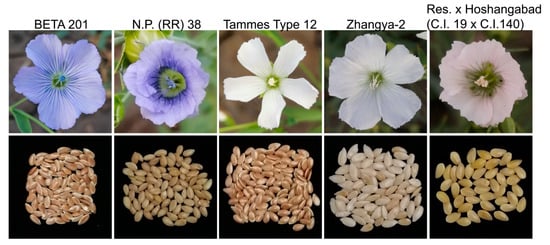

Figure 1
Open AccessArticle
Biogeographical Ancestry Analyses Using the ForenSeqTM DNA Signature Prep Kit and Multiple Prediction Tools
by
Nina Mjølsnes Salvo, Gunn-Hege Olsen, Thomas Berg and Kirstin Janssen
Genes 2024, 15(4), 510; https://doi.org/10.3390/genes15040510 - 18 Apr 2024
Abstract
The inference of biogeographical ancestry (BGA) can assist in police investigations of serious crime cases and help to identify missing people and victims of mass disasters. In this study, we evaluated the typing performance of 56 ancestry-informative SNPs in 177 samples using the
[...] Read more.
The inference of biogeographical ancestry (BGA) can assist in police investigations of serious crime cases and help to identify missing people and victims of mass disasters. In this study, we evaluated the typing performance of 56 ancestry-informative SNPs in 177 samples using the ForenSeq™ DNA Signature Prep Kit on the MiSeq FGx system. Furthermore, we compared the prediction accuracy of the tools Universal Analysis Software v1.2 (UAS), the FROG-kb, and GenoGeographer when inferring the ancestry of 503 Europeans, 22 non-Europeans, and 5 individuals with co-ancestry. The kit was highly sensitive with complete aiSNP profiles in samples with as low as 250pg input DNA. However, in line with others, we observed low read depth and occasional drop-out in some SNPs. Therefore, we suggest not using less than the recommended 1ng of input DNA. FROG-kb and GenoGeographer accurately predicted both Europeans (99.6% and 91.8% correct, respectively) and non-Europeans (95.4% and 90.9% correct, respectively). The UAS was highly accurate when predicting Europeans (96.0% correct) but performed poorer when predicting non-Europeans (40.9% correct). None of the tools were able to correctly predict individuals with co-ancestry. Our study demonstrates that the use of multiple prediction tools will increase the prediction accuracy of BGA inference in forensic casework.
Full article
(This article belongs to the Special Issue State-of-the-Art in Forensic Genetics Volume II)
►▼
Show Figures
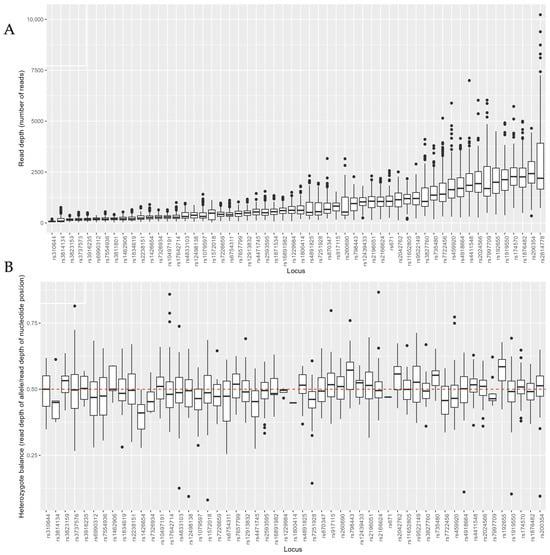
Figure 1

Journal Menu
► ▼ Journal Menu-
- Genes Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biology, BioMed, Cells, Genes
Applications of the Zebrafish Model
Topic Editors: De-Li Shi, Pengfei XuDeadline: 30 June 2024
Topic in
Biomedicines, Cells, CIMB, Genes, IJMS
Animal Models of Human Disease 2.0
Topic Editors: Sigrun Lange, Jameel M. InalDeadline: 31 August 2024
Topic in
Agriculture, Bioengineering, Genes, IJMS, Plants
Genetic Engineering in Agriculture
Topic Editors: Amy L. Klocko, Jianjun Chen, Haiwei LuDeadline: 30 September 2024
Topic in
Biology, BioMedInformatics, Cancers, Genes, IJMS
The 22nd International Conference on Bioinformatics (InCoB 2023): Translational Bioinformatics Transforming Life
Topic Editors: Jyotsna Batra, Srilakshmi Srinivasan, Shoba Ranganathan, Asif M. Khan, Harpreet SinghDeadline: 15 November 2024

Conferences
Special Issues
Special Issue in
Genes
Molecular Mechanisms Responsible for Radiation-Induced Toxicity of Normal Tissue
Guest Editor: Rupak PathakDeadline: 25 April 2024
Special Issue in
Genes
Cotton Genes, Genetics, and Genomics
Guest Editor: Guanjing HuDeadline: 10 May 2024
Special Issue in
Genes
DNA Damage and Repair in Microorganisms, Plants and Mammalian Systems
Guest Editors: Ioly Kotta-Loizou, Nan Zhang, Xin WangDeadline: 20 May 2024
Special Issue in
Genes
Commemorating the Launch of the Section "Cytogenomics"
Guest Editor: Darren GriffinDeadline: 5 June 2024
Topical Collections
Topical Collection in
Genes
Study on Genotypes and Phenotypes of Pediatric Clinical Rare Diseases
Collection Editors: Livia Garavelli, Stefano Giuseppe Caraffi
Topical Collection in
Genes
Eukaryotic Non-coding RNAs: Diversity, Structure/Function, Implication in Cardiovascular Disease
Collection Editors: Morten Andre Høydal, Christiane Branlant
Topical Collection in
Genes
Feature Papers in ‘Animal Genetics and Genomics’
Collection Editors: Antonio Figueras, Raquel Vasconcelos







